Spaces:
Sleeping
title: Qubit-Medic
emoji: 🩺
colorFrom: indigo
colorTo: pink
sdk: docker
app_port: 7860
pinned: true
tags:
- openenv
- reinforcement-learning
- quantum-error-correction
- stim
- pymatching
- grpo
- trl
- llm
license: mit
short_description: OpenEnv RL env that teaches an LLM to decode quantum errors.
Qubit-Medic
An LLM trained to decode quantum surface-code syndromes. We follow the AlphaQubit-style recipe (Nature 2024): a language model as decoder with verifiable rewards—implemented on Stim + PyMatching, an OpenEnv-style HTTP contract, SFT warm-up + GRPO (TRL/Unsloth), and multi-component rewards that are hard to game.
Hugging Face
- Space: https://huggingface.co/spaces/ronitraj/QuantumScribe — live OpenEnv server + API; liveness: https://ronitraj-quantumscribe.hf.space/healthz
- Model (LoRA): https://huggingface.co/ronitraj/quantumscribe — PEFT adapter and tokenizer
Weights & Biases (this experiment)
- Project: https://wandb.ai/ronitraj/QuantumScribe-GRPO
- SFT run (
sft-20260426-045056): https://wandb.ai/ronitraj/QuantumScribe-GRPO/runs/yli513jl - GRPO run (
grpo-20260426-045324): https://wandb.ai/ronitraj/QuantumScribe-GRPO/runs/4p7eurnc — run id4p7eurnc(e.g. best step ~1300, in-loop eval, artifacts)
Quick links
| Resource | URL |
|---|---|
| Hugging Face Space (live demo + API) | ronitraj/QuantumScribe — health: /healthz |
| Trained LoRA on the Hub | ronitraj/quantumscribe (PEFT adapter + tokenizer) |
| W&B project | ronitraj/QuantumScribe-GRPO |
| W&B — SFT run | runs/yli513jl |
| W&B — GRPO run | runs/4p7eurnc |
| Colab training | notebooks/colab_train.ipynb |
| Local Gradio | python app_gradio.py |
| OpenEnv manifest | openenv.yaml |
What this repo does (elevator pitch)
Quantum computers need a decoder: classical software that maps syndromes (detector results) to corrections. DeepMind’s AlphaQubit showed a transformer can beat a strong PyMatching baseline. We reimplement the idea with a commodity stack:
- 3B instruction-tuned Qwen2.5 in 4-bit (Unsloth) + LoRA
- SFT then GRPO (reward from a real Stim environment, not offline labels)
- OpenEnv-compatible server:
/reset//step/ state & schema - Five logged reward components (aggregate is weighted)
| Dimension | This project (typical) | AlphaQubit (reference) |
|---|---|---|
| Decoder | 3B LM + LoRA (off-the-shelf) | Custom architecture, lab-scale data mix |
| Training signal | SFT + GRPO on env reward | Proprietary + SI1000 / Sycamore |
| Baseline | PyMatching (sparse blossom) | Same class of MWM decoder |
| Open source | This repo + Hub weights | Research partial |
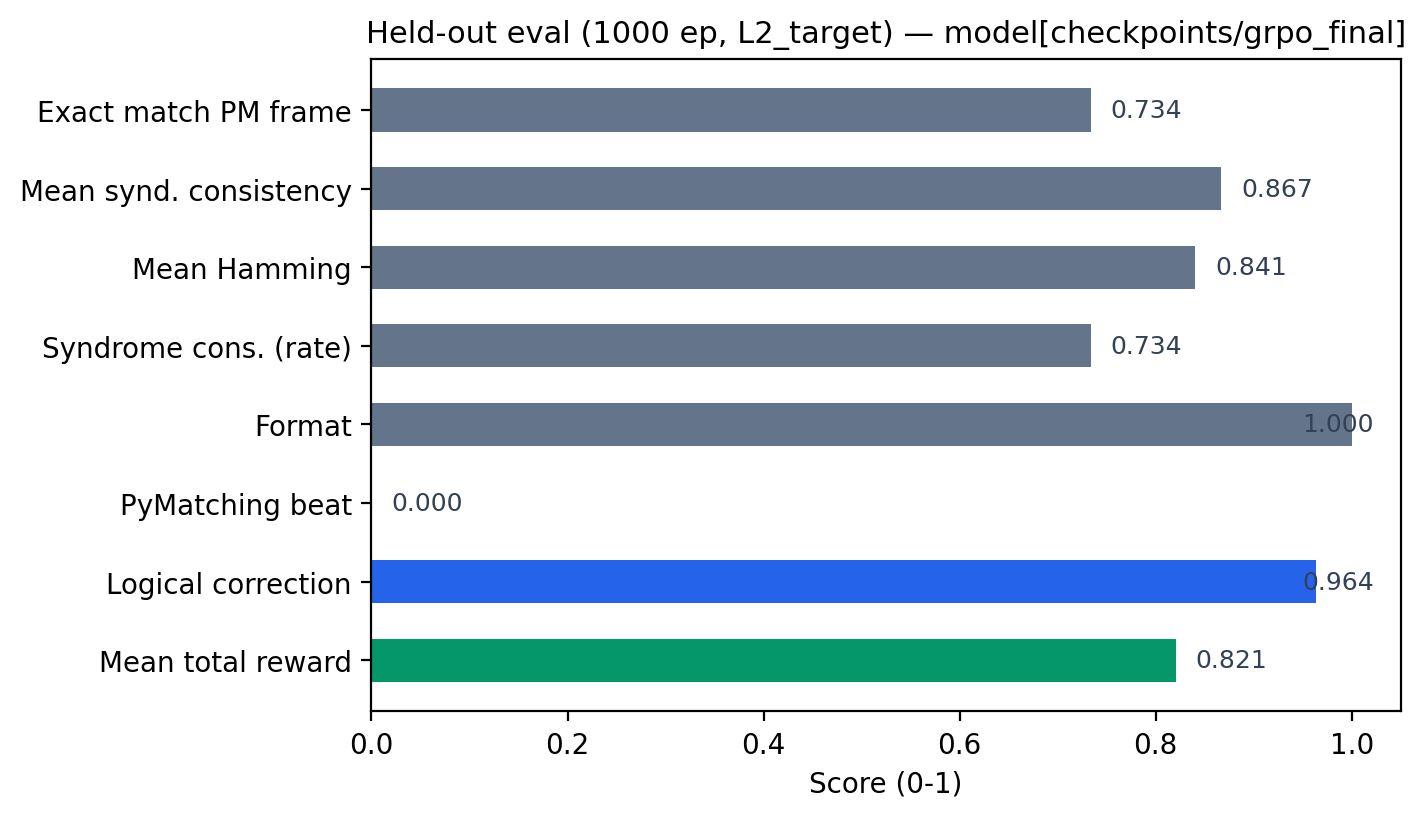
Latest measured eval (JSON)
These numbers come from a held-out run written to data/eval_grpo.json (1000 episodes, L2 target, adapter path recorded in the file). They are the source of truth for submission claims; do not substitute synthetic plots for these metrics.
| Metric | Value |
|---|---|
logical_correction_rate |
0.964 |
pymatching_beat_rate |
0.0 |
format_compliance_rate |
1.0 |
mean_hamming_overlap |
0.8405 |
mean_total_reward |
~0.821 |
exact_match_pymatching |
0.734 |
pymatching_beat is 1 only when PyMatching is wrong on the observable and the LLM is right; on this eval it is 0.0—i.e. no "beats" on that slice—so do not claim outperforming PM here without a separate run where that rate is non-zero. High logical correction and overlap with the PM frame remain meaningful; interpret with reward definitions.
Reproduce:
python -m scripts.eval --adapter /path/to/grpo/adapter --episodes 1000 --out data/eval_grpo.json
(Adjust --adapter to your checkpoint, e.g. a downloaded ronitraj/quantumscribe adapter.)
Data in data/
| File | Purpose |
|---|---|
| data/eval_grpo.json | Primary eval — single JSON summary (episodes, logical_correction_rate, pymatching_beat_rate, overlaps, level, etc.) from scripts.eval. |
| data/grpo_validation.jsonl | GRPO validation prompts / episodes (one JSON object per line; curriculum, syndrome, seeds). |
| data/sft_dataset_analysis.json | SFT dataset report — stats (completion lengths, level mix, train/val overlap, eval_windows). |
| data/sft_validation.jsonl | SFT held-out set used during training. |
| data/sft_dataset_sample.jsonl | Small sample of SFT training rows (prompt + metadata). |
Generated on demand (not always committed) after make baselines / SFT / Willow runs, per .gitignore:
data/baseline_results.json— random / zeros / PyMatching baselinesdata/sft_dataset.jsonl— full SFT train (frommake sft-dataorgenerate_sft_data)data/willow_validation.json,data/willow_d3.dem— cross-distribution checks
Figures in figures/
Provenance and regeneration: figures/FIGURES.md. The three trajectory plots below are illustrative (from make plots / baseline-anchored synthetic mode), not a raw W&B export—replace with scripts/plot_results.py and real logs when you have them.
Training trajectories (illustrative)
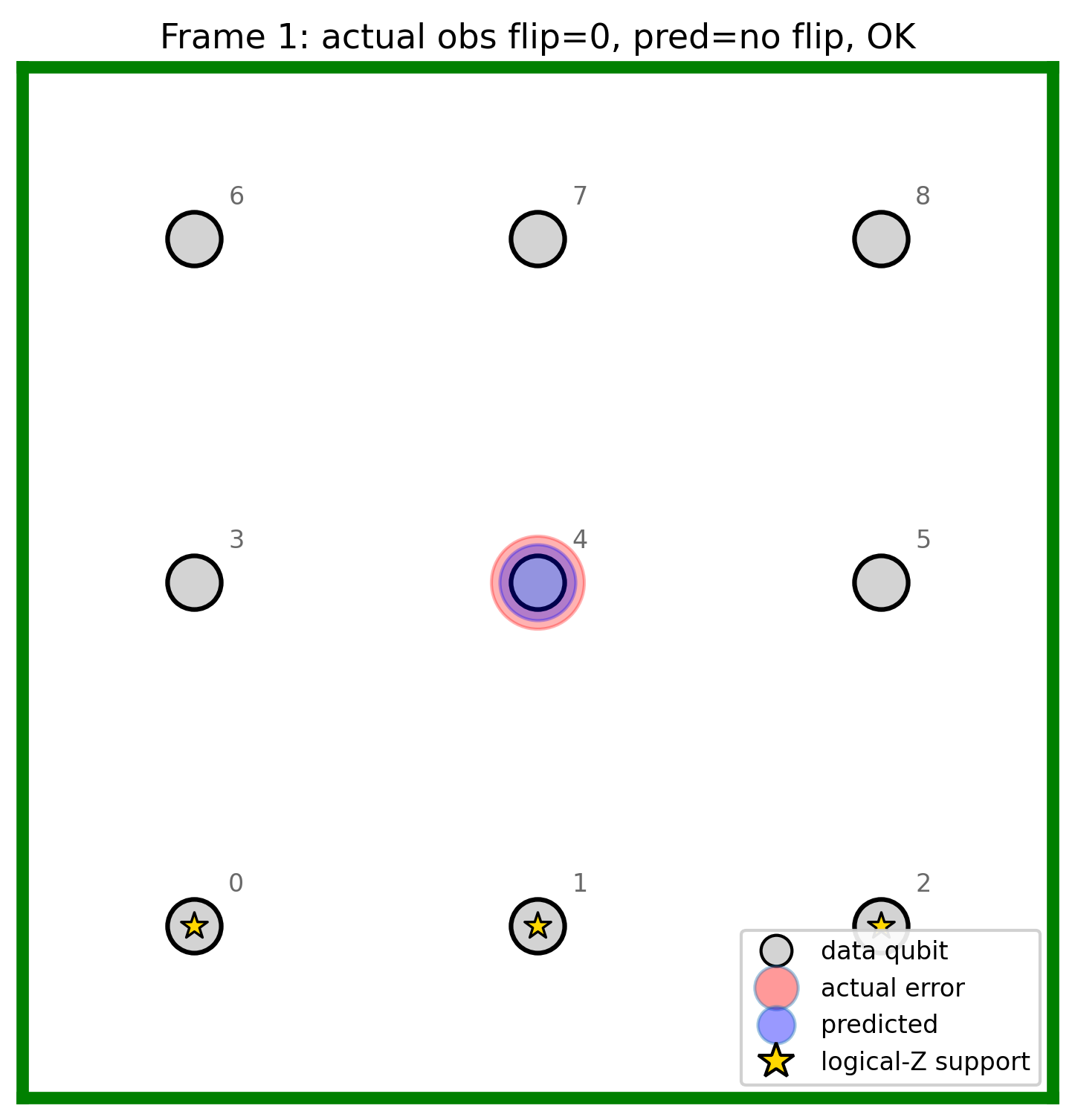
Grid animation (Stim + layout demo)
Reward & metrics from data (reproducible) — not time-series; single-run summaries from data/eval_grpo.json and data/sft_dataset_analysis.json. Regenerate: python -m scripts.plot_data_figures
Note: For per-reward time series and KL during GRPO, use the main GRPO run: runs/4p7eurnc — e.g. rl/reward/total_mean, rl/reward/logical_correction_mean, alarms/kl_alarm_value.
The problem (in one story)
Qubits are noisy. You do not observe errors directly; you get syndromes from stabilizer measurements. A decoder turns syndromes into a Pauli correction. PyMatching is a strong classical baseline. We train an LLM to output a parseable correction; the environment checks it with Stim and five reward functions.
The environment
A FastAPI app exposes an OpenEnv-style flow (see qubit_medic/server/app.py and qubit_medic/server/openenv_adapter.py):
reset(seed)— sample a syndrome (curriculum), return a prompt.step(text)— parse, score rewards, return reward + per-componentinfo.
Episodes are single-step: one completion per episode. The trainer and W&B see each reward component separately.
+----------+ reset / step +---------------------------+
| TRL/ | ------------> | Qubit-Medic (Stim+PM) |
| Unsloth | observation | parse, 5 rewards, return |
+----------+ <------------ +---------------------------+
Methodology checklist
| Concern | Status | Pointer |
|---|---|---|
| Realistic noise (SI1000) | Used | Gidney & Fowler arXiv:2108.10457 |
| Real code family | Stim surface_code:rotated_memory_z |
Stim |
| Strong classical baseline | PyMatching v2 | arXiv:2303.15933 |
| Policy optimisation | GRPO | arXiv:2402.03300 |
| OOD / Willow (optional) | scripts/willow_validation.py + data/willow_d3.dem |
Zenodo |
Baselines (no LLM)
make baselines writes data/baseline_results.json (random, all-zeros, PyMatching). make plots rebuilds the headline figures from that JSON (see figures/FIGURES.md).
make baselines
make plots
Reward design (config-driven)
Weights are qubit_medic/config.py → REWARD_WEIGHTS (sum 1.0):
total = 0.35 * logical_correction
+ 0.25 * hamming_overlap
+ 0.20 * syndrome_consistency
+ 0.10 * format_compliance
+ 0.10 * pymatching_beat
| Component | Role |
|---|---|
| logical_correction | 1 if the implied correction matches logical observable (Stim). |
| hamming_overlap | Dense credit vs the PyMatching reference frame. |
| syndrome_consistency | Implied final detectors vs observed syndrome. |
| format_compliance | Parse success / partial / fail. |
| pymatching_beat | 1 only if PM wrong and LLM right (rare; headline for beating PM). |
Details: qubit_medic/server/rewards.py. GRPO uses a shared batch cache so all five components score the same (prompt, completion) (see W&B section in previous docs—qubit_medic/wandb_utils.py and trainer).
Weights & Biases
Defaults: WANDB_ENTITY=ronitraj, WANDB_PROJECT=QuantumScribe-GRPO. Trainers use qubit_medic/wandb_utils.py. Disable: WANDB_DISABLED=1 or QUBIT_MEDIC_WANDB=0.
Reference runs (2026-04-26, Colab / server)
| Stage | Run name | Direct link |
|---|---|---|
| Project | — | wandb.ai/ronitraj/QuantumScribe-GRPO |
| SFT | sft-20260426-045056 |
runs/yli513jl |
| GRPO | grpo-20260426-045324 |
runs/4p7eurnc |
The GRPO run includes training curves, in-loop eval/*, alarms/kl_alarm_value, best checkpoint metadata (best/step ≈ 1300), and logged artifacts.
pip install -r requirements-train.txt
wandb login
GROUP=my-exp make train-sft
GROUP=my-exp make train-grpo
GROUP=my-exp make eval
Reproducibility (qubit_medic/config.py)
| Item | Value |
|---|---|
| Stim / PyMatching | Pinned in requirements*.txt |
| SFT default base | Qwen/Qwen2.5-3B-Instruct via Unsloth |
| GRPO default base | unsloth/qwen2.5-3b-instruct-unsloth-bnb-4bit |
| LoRA | r=16, alpha=32, dropout=0.1, q/k/v/o |
| GRPO | 1500 steps, short completions (max_completion 50), KL coeff 0.02, temperature=1.2 rollouts, etc. |
| Seeds | 42, 1337, 2024 |
**Import from qubit_medic.config**—do not duplicate magic numbers in scripts.
Train and eval (local)
python3 -m venv .venv && . .venv/bin/activate
pip install -r requirements.txt
make validate
make sft-data
make baselines
make tests
python -m scripts.train_sft --output checkpoints/sft_warmup
python -m scripts.train_grpo \
--sft-checkpoint checkpoints/sft_warmup/checkpoint-50 \
--output checkpoints/grpo
python -m scripts.eval --adapter checkpoints/grpo --episodes 1000 --out data/eval_grpo.json
End-to-end: notebooks/colab_train.ipynb. Makefile shortcuts: make train-sft, make train-grpo, make eval (see Makefile).
Local dev: run everything (no Docker)
1. Base environment (CPU OK) — OpenEnv / Stim / tests:
cd /path/to/errorCorrection
python3 -m venv .venv
source .venv/bin/activate # Windows: .venv\Scripts\activate
pip install -U pip
pip install -r requirements.txt
make validate
make tests
2. OpenEnv HTTP server (no LLM — physics + reward only) — good for API checks and curl / a browser:
# default: 0.0.0.0:7860 (or set QUBIT_MEDIC_PORT)
python -m qubit_medic.server.app
# dev reload:
uvicorn qubit_medic.server.app:app --reload --host 0.0.0.0 --port 7860
- Docs: http://127.0.0.1:7860/docs
- Health: http://127.0.0.1:7860/healthz
3. Gradio grid demo (Stim + PyMatching only) — does not load the trained LLM in code today; it visualises the classical decoder.
pip install "gradio>=4"
PORT=7860 python app_gradio.py
# open http://127.0.0.1:7860 — if the OpenEnv server is already on 7860, use e.g. PORT=7861
4. Run with the real model (Unsloth + LoRA) — this is the supported path — needs a GPU and training deps. The eval harness loads the adapter and uses LocalDecoderClient (in-process env, no separate server).
pip install -r requirements-train.txt
# optional: export HF_TOKEN=... for gated/private Hub repos
python -m scripts.eval \
--adapter ronitraj/quantumscribe \
--episodes 50 \
--level L2_target \
--max-new-tokens 160
- Use a local LoRA folder the same way:
--adapter /path/to/checkpoints/grpo/final(the directory that containsadapter_model.safetensors). - The script calls
FastLanguageModel.from_pretrained(model_name=adapter, …); for Hub PEFT repos, Unsloth/transformers should resolve the base fromadapter_config.json. If loading fails, runhf download ronitraj/quantumscribeand point--adapterat the local folder. - Shorter run first (e.g.
--episodes 5) to confirm VRAM, then increase.
5. What is not wired — the Docker Space image does not install torch/Unsloth; the Gradio app’s markdown mentions QUBIT_MEDIC_ADAPTER but there is no LLM inference in app_gradio.py yet—use scripts.eval for the trained policy.
Publish the adapter to the Hub
Released weights: ronitraj/quantumscribe. Load as PEFT on the same base used for training:
from peft import PeftModel
from transformers import AutoModelForCausalLM, AutoTokenizer
base = "unsloth/qwen2.5-3b-instruct-unsloth-bnb-4bit"
model = AutoModelForCausalLM.from_pretrained(base, device_map="auto", trust_remote_code=True)
model = PeftModel.from_pretrained(model, "ronitraj/quantumscribe")
tokenizer = AutoTokenizer.from_pretrained("ronitraj/quantumscribe")
Re-upload: hf upload ronitraj/quantumscribe /path/to/final . with Hub authentication.
Space deployment
- Space: ronitraj/QuantumScribe
- Script:
python -m scripts.deploy_to_space— see scripts/deploy_to_space.py - For private model pulls, set Space secret
HF_TOKEN.
Cross-distribution (optional)
python -m scripts.willow_validation — see scripts/willow_validation.py.
Repository layout
qubit_medic/
config.py, models.py, prompts.py, wandb_utils.py
client/
server/ (app, environment, rewards, curriculum, physics, openenv_adapter)
scripts/
validate_env.py, generate_sft_data.py, train_sft.py, train_grpo.py, eval.py
baseline_policies.py, plot_results.py, plot_data_figures.py, animate_grid.py, willow_validation.py
format_test.py, diversity_preflight.py, deploy_to_space.py, sync_kaggle_bundle.py
tests/ data/ figures/ checkpoints/ notebooks/colab_train.ipynb
app_gradio.py Dockerfile openenv.yaml Makefile
Citations
@article{bausch_alphaqubit_2024,
title = {Learning high-accuracy error decoding for quantum processors},
author = {Bausch, Johannes and others},
journal = {Nature},
volume = {635},
pages = {834},
year = {2024},
doi = {10.1038/s41586-024-08148-8}
}
@article{acharya_willow_2024,
title = {Quantum error correction below the surface code threshold},
author = {Acharya, R. and others (Google Quantum AI)},
journal = {arXiv:2408.13687},
year = {2024}
}
@article{gidney_si1000_2021,
title = {A fault-tolerant honeycomb memory},
author = {Gidney, Craig and Fowler, Austin G.},
journal = {arXiv:2108.10457},
year = {2021}
}
@article{higgott_pymatching_2023,
title = {Sparse Blossom: correcting a million errors per core second
with minimum-weight matching},
author = {Higgott, Oscar and Gidney, Craig},
journal = {arXiv:2303.15933},
year = {2023}
}
@article{shao_grpo_2024,
title = {DeepSeekMath: pushing the limits of mathematical reasoning
in open language models},
author = {Shao, Zhihong and others},
journal = {arXiv:2402.03300},
year = {2024}
}
Acknowledgments
DeepMind (AlphaQubit), Google Quantum AI (Stim, Willow data), Gidney (SI1000), Higgott (PyMatching), Hugging Face, Unsloth, OpenEnv.
License
MIT — LICENSE.