TriALS: Triphasic-Aided Liver Lesion Segmentation Benchmark in Non-Contrast CT
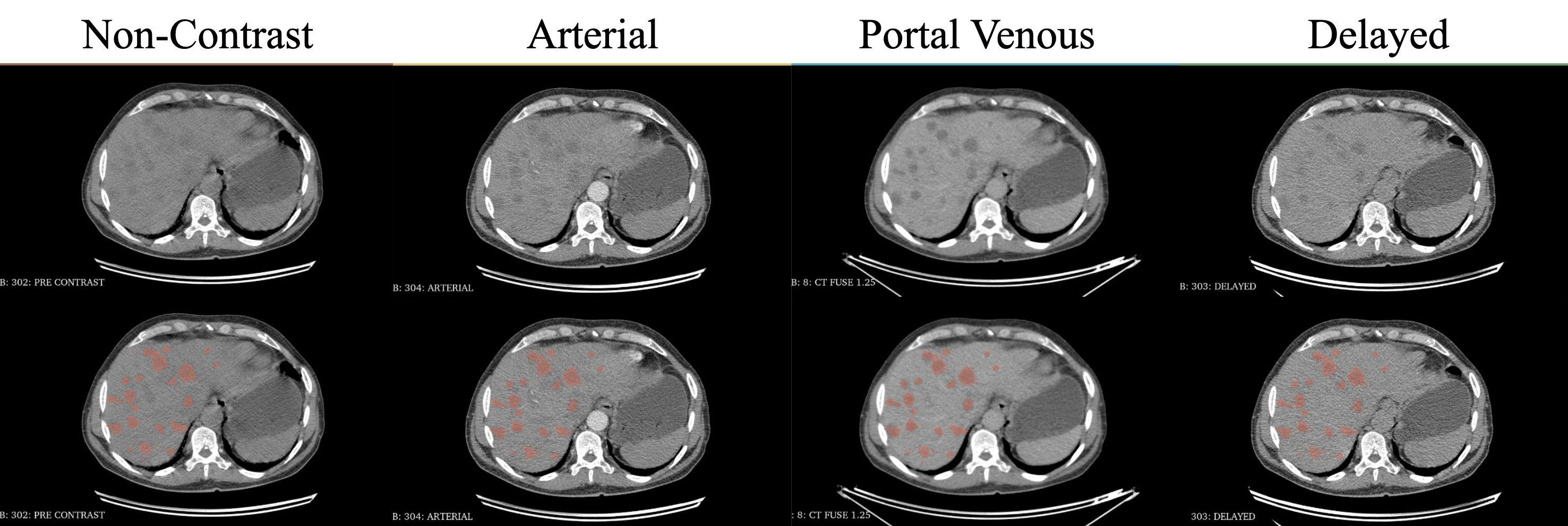
Same patient across all four CT phases (non-contrast, arterial, portal venous, delayed). Top row: raw images. Bottom row: lesion annotations. Many lesions are occult on non-contrast CT and only become conspicuous after contrast administration — this is the core diagnostic challenge TriALS targets.
TriALS is the first multi-centre benchmark for liver lesion segmentation in non-contrast CT (NCCT), supported by aligned four-phase acquisitions (non-contrast, arterial, portal venous, delayed) from Egyptian and Chinese institutions. It was organised across the MICCAI 2024 and 2025 challenges and is designed to enable development of diagnostic AI for contrast-limited clinical settings.
- Paper: TriALS: Triphasic-Aided Liver Lesion Segmentation Benchmark in Non-Contrast CT (2026)
- Code & evaluation: https://github.com/xmed-lab/TriALS
- License: CC BY 4.0
Dataset summary
| Released cases | 80 (training only) |
| Volumes released | 320 (4 phases × 80) |
| Institutions | Ain Shams University (Egypt), Sun Yat-Sen Memorial Hospital (China) |
| Modality | Abdominal CT, NIfTI, native clinical resolution, Hounsfield units (no resampling) |
| Annotations | Per-phase lesion masks (4) + combined label registered to NCCT |
Note on the test sets. Egypt internal, China internal, and the external (Nanfang Hospital) cohorts are not released publicly and are retained for ongoing validation. Researchers who wish to evaluate their method on these cohorts can contact the corresponding authors to arrange held-out evaluation via the challenge pipeline.
Tasks
- Task 1 — Venous-phase segmentation. Standard contrast-enhanced lesion segmentation on the portal venous phase.
- Task 2 — Non-contrast segmentation. Lesion segmentation on NCCT under two paradigms:
- Visible: evaluated against lesions annotated from NCCT alone.
- Combined: evaluated against the multi-phase fused label (full lesion burden, including lesions occult on NCCT). This is the primary clinical target.
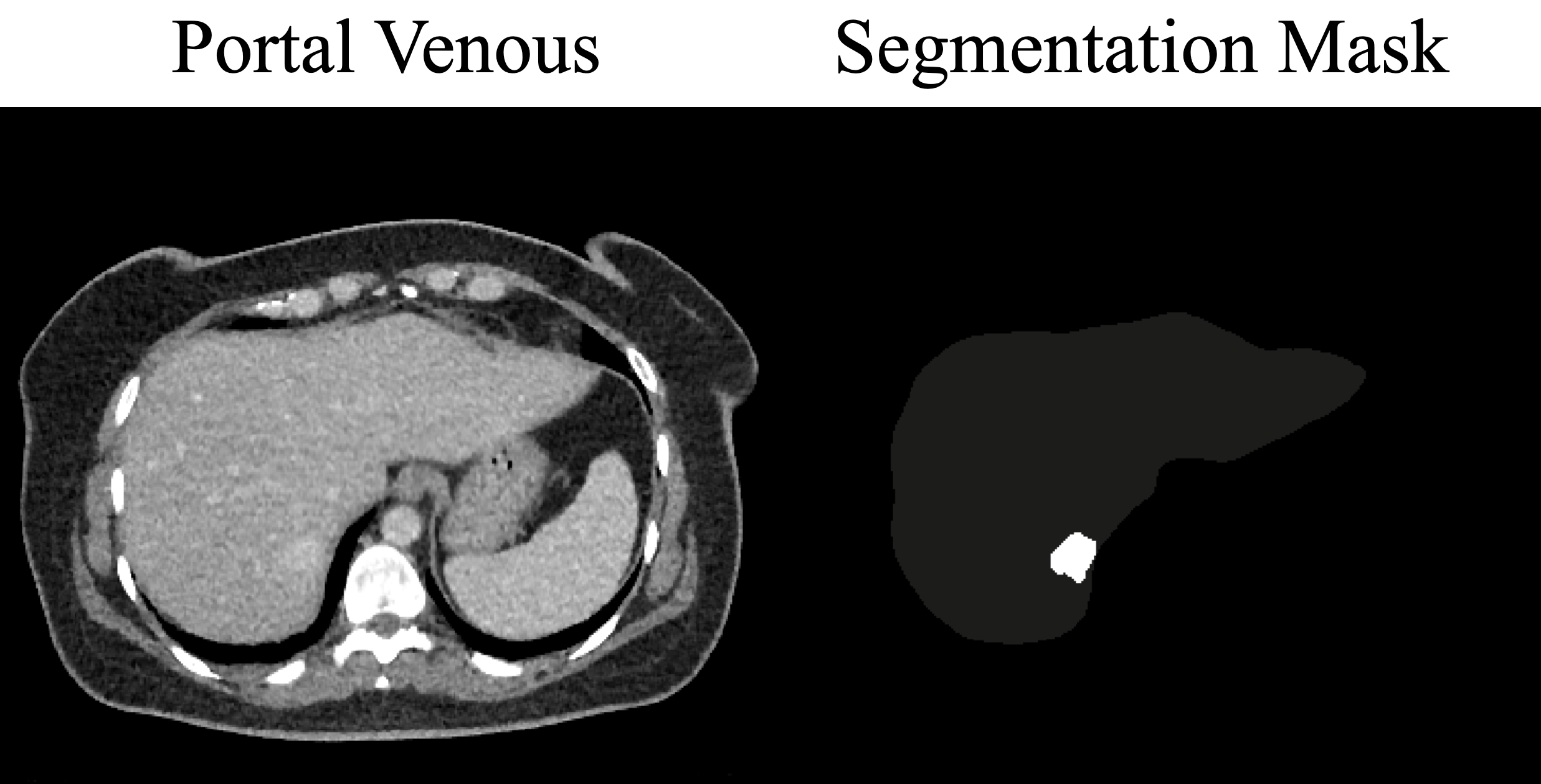
A single case: portal venous CT (left) and the corresponding lesion label within the liver region (right).
Directory structure
Dataset_TriALS/
├── imagesTr/
│ ├── TriALS-0/
│ │ ├── TriALS-0_nocontrast.nii.gz
│ │ ├── TriALS-0_arterial.nii.gz
│ │ ├── TriALS-0_venous.nii.gz
│ │ └── TriALS-0_delayed.nii.gz
│ ├── TriALS-1/
│ │ └── ...
│ └── TriALS-<caseID>/
└── labelsTr/
├── TriALS-0/
│ ├── TriALS-0_nocontrast.nii.gz
│ ├── TriALS-0_arterial.nii.gz
│ ├── TriALS-0_venous.nii.gz
│ ├── TriALS-0_delayed.nii.gz
│ └── TriALS-0_combined.nii.gz
├── TriALS-1/
│ └── ...
└── TriALS-<caseID>/
Each case has its own folder under imagesTr/ and labelsTr/. Filenames follow TriALS-<caseID>_<phase>.nii.gz.
Case ID convention
| Centre | Case ID range | n |
|---|---|---|
| Egypt (Ain Shams University) | TriALS-0 … TriALS-59 |
60 |
| China (Sun Yat-Sen Memorial) | TriALS-200 … TriALS-219 |
20 |
China cases are numbered from 200 onwards to keep centre identity recoverable from the filename.
Phase convention
| Phase tag | Meaning | In images | In labels |
|---|---|---|---|
nocontrast |
Non-contrast CT | ✓ | ✓ |
arterial |
Arterial phase | ✓ | ✓ |
venous |
Portal venous phase | ✓ | ✓ |
delayed |
Delayed phase | ✓ | ✓ |
combined |
Multi-phase fused label, registered to NCCT | — | ✓ |
Download
Full dataset
from huggingface_hub import snapshot_download
path = snapshot_download(
repo_id="marwankefah/TriALS",
repo_type="dataset",
)
Selective download by task and centre
Choose TASK ∈ {"task1", "task2", "all"} and CENTRE ∈ {"egypt", "china", "both"}:
from huggingface_hub import snapshot_download
# ---- Configure ----
TASK = "task1" # "task1" (venous), "task2" (NCCT + combined label), or "all"
CENTRE = "both" # "egypt", "china", or "both"
# -------------------
centre_ranges = {
"egypt": list(range(0, 60)),
"china": list(range(200, 220)),
"both": list(range(0, 60)) + list(range(200, 220)),
}
task_phases = {
"task1": {"images": ["venous"],
"labels": ["venous"]},
"task2": {"images": ["nocontrast"],
"labels": ["nocontrast", "combined"]},
"all": {"images": ["nocontrast", "arterial", "venous", "delayed"],
"labels": ["nocontrast", "arterial", "venous", "delayed", "combined"]},
}
phases = task_phases[TASK]
patterns = []
for cid in centre_ranges[CENTRE]:
case = f"TriALS-{cid}"
for p in phases["images"]:
patterns.append(f"Dataset_TriALS/imagesTr/{case}/{case}_{p}.nii.gz")
for p in phases["labels"]:
patterns.append(f"Dataset_TriALS/labelsTr/{case}/{case}_{p}.nii.gz")
path = snapshot_download(
repo_id="marwankefah/TriALS",
repo_type="dataset",
allow_patterns=patterns,
)
print(f"Downloaded to: {path}")
A few common recipes:
# Task 1, Egyptian cohort only
TASK, CENTRE = "task1", "egypt"
# Task 2 with multi-phase training data (all 4 image phases + NCCT/combined labels)
TASK, CENTRE = "all", "both"
# then restrict labels in your dataloader to ["nocontrast", "combined"]
Load a volume
import os, nibabel as nib
root = f"{path}/Dataset_TriALS"
case = "TriALS-0"
phase = "venous"
img = nib.load(os.path.join(root, "imagesTr", case, f"{case}_{phase}.nii.gz"))
lbl = nib.load(os.path.join(root, "labelsTr", case, f"{case}_{phase}.nii.gz"))
# Combined multi-phase label (labels only, registered to NCCT)
combined = nib.load(os.path.join(root, "labelsTr", case, f"{case}_combined.nii.gz"))
Annotation protocol
Two-stage hybrid human–algorithm protocol. An automated model trained on LiTS and pilot in-house cases produced preliminary segmentations, which were propagated across phases via non-rigid registration. These were then manually corrected by 10 trained annotators (radiology residents/fellows) in 3D Slicer; each case was reviewed by a second annotator, with disagreements escalated to a senior abdominal radiology consultant.
Combined label
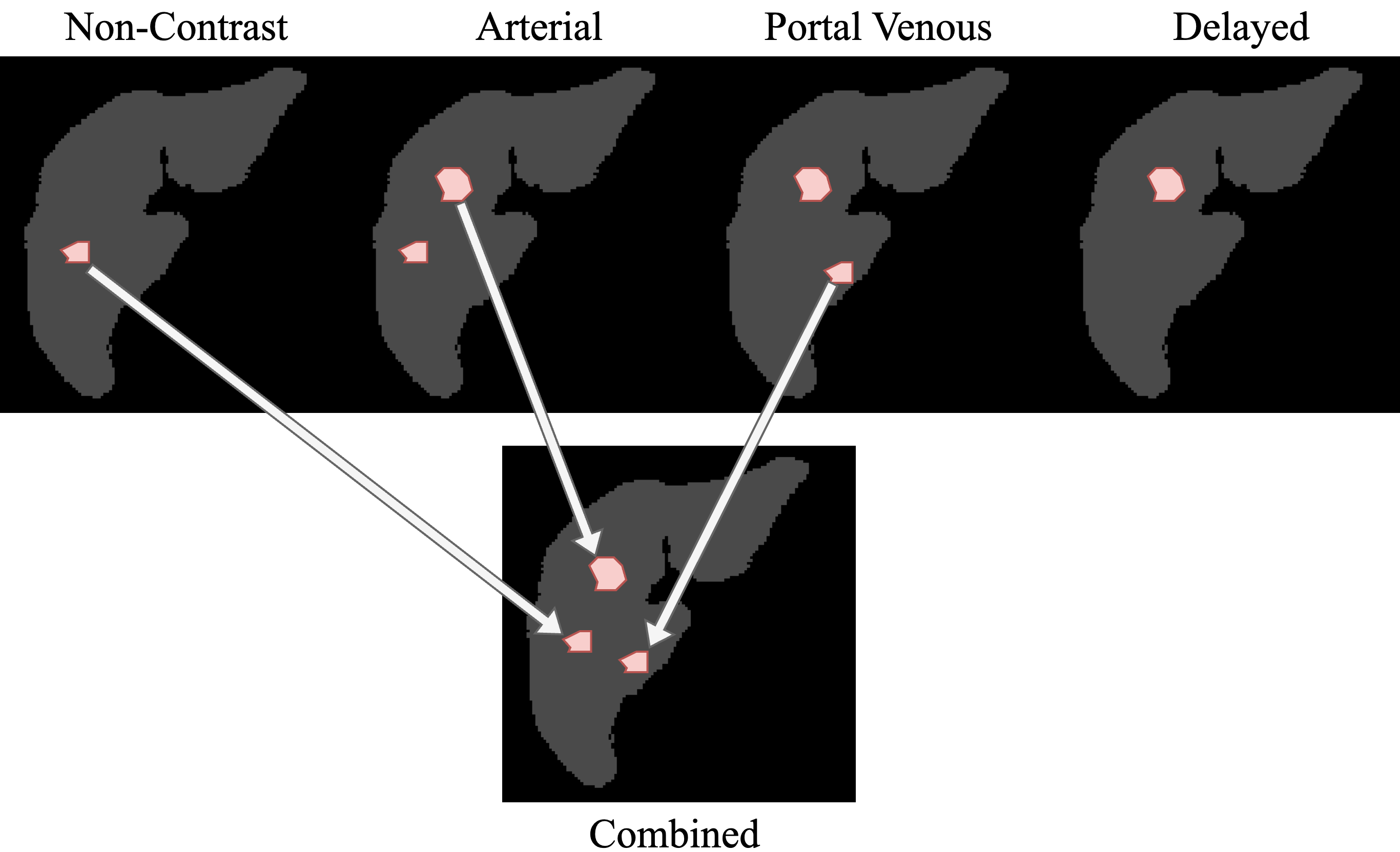
Lesions annotated on each contrast phase (non-contrast, arterial, venous, delayed) are registered to the non-contrast frame and fused into a single combined label that captures the full lesion burden — including lesions occult on NCCT alone.
The combined label was constructed by rigid-then-non-rigid registration of the contrast phases to NCCT via elastix, followed by multi-phase label fusion and consultant-level QA. All raw per-phase volumes and per-phase labels are released alongside the combined labels to enable independent verification of the registration and fusion pipeline.
Ethics and data governance
- Egyptian cohort (Ain Shams University Hospitals, Cairo): collected under local Research Ethics Committee approval (FWA 000017585), with waiver of individual informed consent for retrospective de-identified use.
- Chinese cohort (Sun Yat-Sen Memorial Hospital, Sun Yat-Sen University): approved by the institutional review committee of Sun Yat-sen Memorial Hospital, Sun Yat-sen University.
All volumes were fully anonymised prior to transfer; direct patient identifiers, dates, and institution-identifying metadata were removed or shifted.
Citation
If you use TriALS, please cite:
Acknowledgements
Supported by the Research Grants Council of Hong Kong (T45-401/22-N), the National Natural Science Foundation of China (62306254; 82001768), and the Guangdong Basic and Applied Basic Research Foundation (2021A1515010226). TriALS is endorsed by SIG-AFRICAI (MICCAI Society).
Contact
For questions, collaboration, or requests to evaluate on the held-out test cohorts, contact the corresponding authors listed on the paper.
- Downloads last month
- 18