license: bsd-3-clause
library_name: braindecode
pipeline_tag: feature-extraction
tags:
- eeg
- biosignal
- pytorch
- neuroscience
- braindecode
- foundation-model
- transformer
CBraMod
C\ riss-\ C\ ross Bra\ in Mod\ el for EEG Decoding from Wang et al. (2025) .
Architecture-only repository. This repo documents the
braindecode.models.CBraModclass. No pretrained weights are distributed here — instantiate the model and train it on your own data, or fine-tune from a published foundation-model checkpoint separately.
Quick start
pip install braindecode
from braindecode.models import CBraMod
model = CBraMod(
n_chans=22,
sfreq=200,
input_window_seconds=4.0,
n_outputs=2,
)
The signal-shape arguments above are example defaults — adjust them to match your recording.
Documentation
- Full API reference (parameters, references, architecture figure): https://braindecode.org/stable/generated/braindecode.models.CBraMod.html
- Interactive browser with live instantiation: https://huggingface.co/spaces/braindecode/model-explorer
- Source on GitHub: https://github.com/braindecode/braindecode/blob/master/braindecode/models/cbramod.py#L23
Architecture description
The block below is the rendered class docstring (parameters, references, architecture figure where available).
Criss-Cross Brain Model for EEG Decoding from Wang et al. (2025) [cbramod].
Foundation ModelAttention/Transformer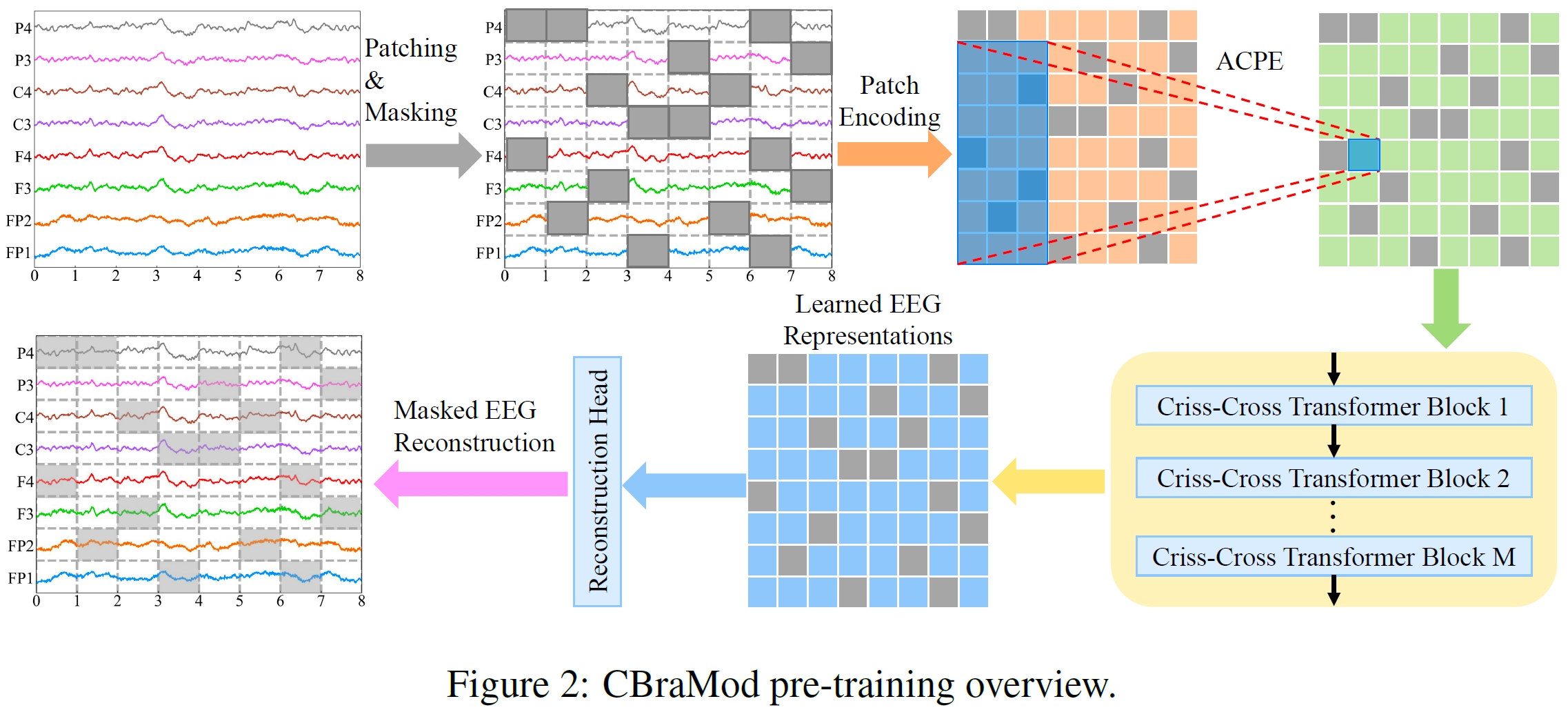
CBraMod is a foundation model for EEG decoding that leverages a novel criss-cross transformer architecture to effectively model the unique spatial and temporal characteristics of EEG signals. Pre-trained on the Temple University Hospital EEG Corpus (TUEG)—the largest public EEG corpus— using masked EEG patch reconstruction, CBraMod achieves state-of-the-art performance across diverse downstream BCI and clinical applications.
Key Innovation: Criss-Cross Attention
Unlike existing EEG foundation models that use full attention to model all spatial and temporal dependencies together, CBraMod separates spatial and temporal dependencies through a criss-cross transformer architecture:
Spatial Attention: Models dependencies between channels while keeping patches separate
Temporal Attention: Models dependencies between temporal patches while keeping channels separate
This design is inspired by criss-cross strategies from computer vision and effectively leverages the inherent structural characteristics of EEG signals. The criss-cross approach reduces computational complexity (FLOPs reduced by ~32% compared to full attention) while improving performance and enabling faster convergence.
Asymmetric Conditional Positional Encoding (ACPE)
Rather than using fixed positional embeddings, CBraMod employs Asymmetric Conditional Positional Encoding that dynamically generates positional embeddings using a convolutional network. This enables the model to:
Capture relative positional information adaptively
Handle diverse EEG channel formats (different channel counts and reference schemes)
Generalize to arbitrary downstream EEG formats without retraining
Support various reference schemes (earlobe, average, REST, bipolar)
Pretraining Highlights
Pretraining Dataset: Temple University Hospital EEG Corpus (TUEG), the largest public EEG corpus
Pretraining Task: Self-supervised masked EEG patch reconstruction from both time-domain and frequency-domain EEG signals
Model Parameters: ~4.0M parameters (very compact compared to other foundation models)
Fast Convergence: Achieves decent results in first epoch on downstream tasks, full convergence within ~10 epochs (vs. ~30 for supervised models like EEGConformer)
Macro Components
Patch Encoding Network: Converts raw EEG patches into embeddings
Asymmetric Conditional Positional Encoding (ACPE): Generates spatial-temporal positional embeddings adaptively from input EEG format
Criss-Cross Transformer Blocks (12 layers): Alternates spatial and temporal attention to learn EEG representations
Reconstruction Head: Reconstructs masked EEG patches during pretraining
- Task head (final_layer): flatten summary tokens across patches and map to
n_outputs; if return_encoder_output=True, return the encoder features instead.
The model is highly efficient, requiring only ~318.9M FLOPs on a typical 16-channel, 10-second EEG recording (significantly lower than full attention baselines).
Known Limitations
Data Quality: TUEG corpus contains "dirty data"; pretraining used crude filtering, reducing available pre-training data
Channel Dependency: Performance degrades with very sparse electrode setups (e.g., <4 channels)
Computational Resources: While efficient, foundation models have higher deployment requirements than lightweight models
Limited Scaling Exploration: Future work should explore scaling laws at billion-parameter levels and integration with large pre-trained vision/language models
Parameters
- patch_sizeint, default=200
Temporal patch size in samples (200 samples = 1 second at 200 Hz).
- dim_feedforwardint, default=800
Dimension of the feedforward network in Transformer layers.
- n_layerint, default=12
Number of Transformer layers.
- nheadint, default=8
Number of attention heads.
- activationtype[nn.Module], default=nn.GELU
Activation function used in Transformer feedforward layers.
- emb_dimint, default=200
Output embedding dimension.
- drop_probfloat, default=0.1
Dropout probability.
- return_encoder_outputbool, default=False
If false (default), the features are flattened and passed through a final linear layer to produce class logits of size n_outputs. If True, the model returns the encoder output features.
References
Wang, J., Zhao, S., Luo, Z., Zhou, Y., Jiang, H., Li, S., Li, T., & Pan, G. (2025). CBraMod: A Criss-Cross Brain Foundation Model for EEG Decoding. In The Thirteenth International Conference on Learning Representations (ICLR 2025). https://arxiv.org/abs/2412.07236
Hugging Face Hub integration
When the optional huggingface_hub package is installed, all models automatically gain the ability to be pushed to and loaded from the Hugging Face Hub. Install with:
pip install braindecode[hub]
Pushing a model to the Hub:
Loading a model from the Hub:
Extracting features and replacing the head:
Saving and restoring full configuration:
All model parameters (both EEG-specific and model-specific such as dropout rates, activation functions, number of filters) are automatically saved to the Hub and restored when loading.
See :ref:`load-pretrained-models` for a complete tutorial.
Citation
Please cite both the original paper for this architecture (see the References section above) and braindecode:
@article{aristimunha2025braindecode,
title = {Braindecode: a deep learning library for raw electrophysiological data},
author = {Aristimunha, Bruno and others},
journal = {Zenodo},
year = {2025},
doi = {10.5281/zenodo.17699192},
}
License
BSD-3-Clause for the model code (matching braindecode). Pretraining-derived weights, if you fine-tune from a checkpoint, inherit the licence of that checkpoint and its training corpus.