Adding files from github
Browse filesThis view is limited to 50 files because it contains too many changes. See raw diff
- CITATION.md +13 -0
- LICENSE +166 -0
- Tutorial/Makefile +177 -0
- Tutorial/Source/GEPPETTO.md +12 -0
- Tutorial/Source/GEPPETTO.ses.js +35 -0
- Tutorial/Source/GEPPETTO.singleap.ses.js +43 -0
- Tutorial/Source/GEPPETTO.vclamp.ses.js +59 -0
- Tutorial/Source/HHCellNetwork.net.nml +24 -0
- Tutorial/Source/HHCellSingleAP.net.nml +36 -0
- Tutorial/Source/HHCellVClamp.net.nml +23 -0
- Tutorial/Source/HodgkinHuxley.py +401 -0
- Tutorial/Source/LEMS_HH_Simulation.xml +69 -0
- Tutorial/Source/LEMS_HH_SingleAP.xml +41 -0
- Tutorial/Source/LEMS_HH_VClamp.xml +67 -0
- Tutorial/Source/__init__.py +3 -0
- Tutorial/Source/hhcell.cell.nml +51 -0
- Tutorial/Source/kChan.channel.nml +22 -0
- Tutorial/Source/naChan.channel.nml +27 -0
- Tutorial/Source/nmllite/HH.py +82 -0
- Tutorial/Source/nmllite/HHTest.gv +7 -0
- Tutorial/Source/nmllite/HHTest.gv.png +0 -0
- Tutorial/Source/nmllite/HHTest.mdf.json +70 -0
- Tutorial/Source/nmllite/HHTest.mdf.yaml +57 -0
- Tutorial/Source/nmllite/HHTest.net.nml +21 -0
- Tutorial/Source/nmllite/HHTest.net.png +0 -0
- Tutorial/Source/nmllite/HHTest.nmllite.json +45 -0
- Tutorial/Source/nmllite/HHTest.nmllite.yaml +31 -0
- Tutorial/Source/nmllite/LEMS_SimHHTest.gv +254 -0
- Tutorial/Source/nmllite/LEMS_SimHHTest.xml +33 -0
- Tutorial/Source/nmllite/SimHHTest.json +13 -0
- Tutorial/Source/nmllite/SimHHTest.yaml +8 -0
- Tutorial/Source/nmllite/hhPop_0.v.dat +2001 -0
- Tutorial/Source/nmllite/hhcell.cell.nml +51 -0
- Tutorial/Source/nmllite/kChan.channel.nml +22 -0
- Tutorial/Source/nmllite/naChan.channel.nml +27 -0
- Tutorial/Source/nmllite/passiveChan.channel.nml +16 -0
- Tutorial/Source/nmllite/report.SimHHTest.txt +8 -0
- Tutorial/Source/passiveChan.channel.nml +16 -0
- Tutorial/Source/run.bat +12 -0
- Tutorial/Source/run.sh +13 -0
- Tutorial/Source/test/.test.jnml.omt +17 -0
- Tutorial/Source/test/.test.jnmlnetpyne.omt +17 -0
- Tutorial/Source/test/.test.jnmlnrn.omt +17 -0
- Tutorial/Source/test/.test.mep +6 -0
- Tutorial/Source/test/.test.python.omt +19 -0
- Tutorial/Source/test/.test.singleap.jnmlbrian2.omt +17 -0
- Tutorial/Source/test/.test.singleap.jnmleden.omt +17 -0
- Tutorial/Source/test/.test.singleap.jnmlmoose.omt +17 -0
- Tutorial/Source/test/.test.singleap.jnmlnrn.omt +17 -0
- Tutorial/Source/test/.test.singleap.jnmlpynnnrn.omt +17 -0
CITATION.md
ADDED
|
@@ -0,0 +1,13 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
## Citing this model
|
| 2 |
+
|
| 3 |
+
The software in this repository is provided under the terms of the [software license](LICENSE) included with it.
|
| 4 |
+
|
| 5 |
+
If you use this model in your research, we respectfully ask you to cite:
|
| 6 |
+
|
| 7 |
+
**The original publication upon which this model is based**
|
| 8 |
+
|
| 9 |
+
- Hodgkin AL, Huxley AF. A quantitative description of membrane current and its application to conduction and excitation in nerve. The Journal of Physiology 1952.
|
| 10 |
+
|
| 11 |
+
**The latest release of this repository**
|
| 12 |
+
|
| 13 |
+
- This link should provide a DOI/citation for the latest version released: https://www.zenodo.org/badge/latestdoi/27819667. If you would like us to make a new release, please [open an issue](../../issues).
|
LICENSE
ADDED
|
@@ -0,0 +1,166 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
GNU LESSER GENERAL PUBLIC LICENSE
|
| 2 |
+
Version 3, 29 June 2007
|
| 3 |
+
|
| 4 |
+
Copyright (C) 2007 Free Software Foundation, Inc. <http://fsf.org/>
|
| 5 |
+
Everyone is permitted to copy and distribute verbatim copies
|
| 6 |
+
of this license document, but changing it is not allowed.
|
| 7 |
+
|
| 8 |
+
|
| 9 |
+
This version of the GNU Lesser General Public License incorporates
|
| 10 |
+
the terms and conditions of version 3 of the GNU General Public
|
| 11 |
+
License, supplemented by the additional permissions listed below.
|
| 12 |
+
|
| 13 |
+
0. Additional Definitions.
|
| 14 |
+
|
| 15 |
+
As used herein, "this License" refers to version 3 of the GNU Lesser
|
| 16 |
+
General Public License, and the "GNU GPL" refers to version 3 of the GNU
|
| 17 |
+
General Public License.
|
| 18 |
+
|
| 19 |
+
"The Library" refers to a covered work governed by this License,
|
| 20 |
+
other than an Application or a Combined Work as defined below.
|
| 21 |
+
|
| 22 |
+
An "Application" is any work that makes use of an interface provided
|
| 23 |
+
by the Library, but which is not otherwise based on the Library.
|
| 24 |
+
Defining a subclass of a class defined by the Library is deemed a mode
|
| 25 |
+
of using an interface provided by the Library.
|
| 26 |
+
|
| 27 |
+
A "Combined Work" is a work produced by combining or linking an
|
| 28 |
+
Application with the Library. The particular version of the Library
|
| 29 |
+
with which the Combined Work was made is also called the "Linked
|
| 30 |
+
Version".
|
| 31 |
+
|
| 32 |
+
The "Minimal Corresponding Source" for a Combined Work means the
|
| 33 |
+
Corresponding Source for the Combined Work, excluding any source code
|
| 34 |
+
for portions of the Combined Work that, considered in isolation, are
|
| 35 |
+
based on the Application, and not on the Linked Version.
|
| 36 |
+
|
| 37 |
+
The "Corresponding Application Code" for a Combined Work means the
|
| 38 |
+
object code and/or source code for the Application, including any data
|
| 39 |
+
and utility programs needed for reproducing the Combined Work from the
|
| 40 |
+
Application, but excluding the System Libraries of the Combined Work.
|
| 41 |
+
|
| 42 |
+
1. Exception to Section 3 of the GNU GPL.
|
| 43 |
+
|
| 44 |
+
You may convey a covered work under sections 3 and 4 of this License
|
| 45 |
+
without being bound by section 3 of the GNU GPL.
|
| 46 |
+
|
| 47 |
+
2. Conveying Modified Versions.
|
| 48 |
+
|
| 49 |
+
If you modify a copy of the Library, and, in your modifications, a
|
| 50 |
+
facility refers to a function or data to be supplied by an Application
|
| 51 |
+
that uses the facility (other than as an argument passed when the
|
| 52 |
+
facility is invoked), then you may convey a copy of the modified
|
| 53 |
+
version:
|
| 54 |
+
|
| 55 |
+
a) under this License, provided that you make a good faith effort to
|
| 56 |
+
ensure that, in the event an Application does not supply the
|
| 57 |
+
function or data, the facility still operates, and performs
|
| 58 |
+
whatever part of its purpose remains meaningful, or
|
| 59 |
+
|
| 60 |
+
b) under the GNU GPL, with none of the additional permissions of
|
| 61 |
+
this License applicable to that copy.
|
| 62 |
+
|
| 63 |
+
3. Object Code Incorporating Material from Library Header Files.
|
| 64 |
+
|
| 65 |
+
The object code form of an Application may incorporate material from
|
| 66 |
+
a header file that is part of the Library. You may convey such object
|
| 67 |
+
code under terms of your choice, provided that, if the incorporated
|
| 68 |
+
material is not limited to numerical parameters, data structure
|
| 69 |
+
layouts and accessors, or small macros, inline functions and templates
|
| 70 |
+
(ten or fewer lines in length), you do both of the following:
|
| 71 |
+
|
| 72 |
+
a) Give prominent notice with each copy of the object code that the
|
| 73 |
+
Library is used in it and that the Library and its use are
|
| 74 |
+
covered by this License.
|
| 75 |
+
|
| 76 |
+
b) Accompany the object code with a copy of the GNU GPL and this license
|
| 77 |
+
document.
|
| 78 |
+
|
| 79 |
+
4. Combined Works.
|
| 80 |
+
|
| 81 |
+
You may convey a Combined Work under terms of your choice that,
|
| 82 |
+
taken together, effectively do not restrict modification of the
|
| 83 |
+
portions of the Library contained in the Combined Work and reverse
|
| 84 |
+
engineering for debugging such modifications, if you also do each of
|
| 85 |
+
the following:
|
| 86 |
+
|
| 87 |
+
a) Give prominent notice with each copy of the Combined Work that
|
| 88 |
+
the Library is used in it and that the Library and its use are
|
| 89 |
+
covered by this License.
|
| 90 |
+
|
| 91 |
+
b) Accompany the Combined Work with a copy of the GNU GPL and this license
|
| 92 |
+
document.
|
| 93 |
+
|
| 94 |
+
c) For a Combined Work that displays copyright notices during
|
| 95 |
+
execution, include the copyright notice for the Library among
|
| 96 |
+
these notices, as well as a reference directing the user to the
|
| 97 |
+
copies of the GNU GPL and this license document.
|
| 98 |
+
|
| 99 |
+
d) Do one of the following:
|
| 100 |
+
|
| 101 |
+
0) Convey the Minimal Corresponding Source under the terms of this
|
| 102 |
+
License, and the Corresponding Application Code in a form
|
| 103 |
+
suitable for, and under terms that permit, the user to
|
| 104 |
+
recombine or relink the Application with a modified version of
|
| 105 |
+
the Linked Version to produce a modified Combined Work, in the
|
| 106 |
+
manner specified by section 6 of the GNU GPL for conveying
|
| 107 |
+
Corresponding Source.
|
| 108 |
+
|
| 109 |
+
1) Use a suitable shared library mechanism for linking with the
|
| 110 |
+
Library. A suitable mechanism is one that (a) uses at run time
|
| 111 |
+
a copy of the Library already present on the user's computer
|
| 112 |
+
system, and (b) will operate properly with a modified version
|
| 113 |
+
of the Library that is interface-compatible with the Linked
|
| 114 |
+
Version.
|
| 115 |
+
|
| 116 |
+
e) Provide Installation Information, but only if you would otherwise
|
| 117 |
+
be required to provide such information under section 6 of the
|
| 118 |
+
GNU GPL, and only to the extent that such information is
|
| 119 |
+
necessary to install and execute a modified version of the
|
| 120 |
+
Combined Work produced by recombining or relinking the
|
| 121 |
+
Application with a modified version of the Linked Version. (If
|
| 122 |
+
you use option 4d0, the Installation Information must accompany
|
| 123 |
+
the Minimal Corresponding Source and Corresponding Application
|
| 124 |
+
Code. If you use option 4d1, you must provide the Installation
|
| 125 |
+
Information in the manner specified by section 6 of the GNU GPL
|
| 126 |
+
for conveying Corresponding Source.)
|
| 127 |
+
|
| 128 |
+
5. Combined Libraries.
|
| 129 |
+
|
| 130 |
+
You may place library facilities that are a work based on the
|
| 131 |
+
Library side by side in a single library together with other library
|
| 132 |
+
facilities that are not Applications and are not covered by this
|
| 133 |
+
License, and convey such a combined library under terms of your
|
| 134 |
+
choice, if you do both of the following:
|
| 135 |
+
|
| 136 |
+
a) Accompany the combined library with a copy of the same work based
|
| 137 |
+
on the Library, uncombined with any other library facilities,
|
| 138 |
+
conveyed under the terms of this License.
|
| 139 |
+
|
| 140 |
+
b) Give prominent notice with the combined library that part of it
|
| 141 |
+
is a work based on the Library, and explaining where to find the
|
| 142 |
+
accompanying uncombined form of the same work.
|
| 143 |
+
|
| 144 |
+
6. Revised Versions of the GNU Lesser General Public License.
|
| 145 |
+
|
| 146 |
+
The Free Software Foundation may publish revised and/or new versions
|
| 147 |
+
of the GNU Lesser General Public License from time to time. Such new
|
| 148 |
+
versions will be similar in spirit to the present version, but may
|
| 149 |
+
differ in detail to address new problems or concerns.
|
| 150 |
+
|
| 151 |
+
Each version is given a distinguishing version number. If the
|
| 152 |
+
Library as you received it specifies that a certain numbered version
|
| 153 |
+
of the GNU Lesser General Public License "or any later version"
|
| 154 |
+
applies to it, you have the option of following the terms and
|
| 155 |
+
conditions either of that published version or of any later version
|
| 156 |
+
published by the Free Software Foundation. If the Library as you
|
| 157 |
+
received it does not specify a version number of the GNU Lesser
|
| 158 |
+
General Public License, you may choose any version of the GNU Lesser
|
| 159 |
+
General Public License ever published by the Free Software Foundation.
|
| 160 |
+
|
| 161 |
+
If the Library as you received it specifies that a proxy can decide
|
| 162 |
+
whether future versions of the GNU Lesser General Public License shall
|
| 163 |
+
apply, that proxy's public statement of acceptance of any version is
|
| 164 |
+
permanent authorization for you to choose that version for the
|
| 165 |
+
Library.
|
| 166 |
+
|
Tutorial/Makefile
ADDED
|
@@ -0,0 +1,177 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Makefile for Sphinx documentation
|
| 2 |
+
#
|
| 3 |
+
|
| 4 |
+
# You can set these variables from the command line.
|
| 5 |
+
SPHINXOPTS =
|
| 6 |
+
SPHINXBUILD = sphinx-build
|
| 7 |
+
PAPER =
|
| 8 |
+
BUILDDIR = _build
|
| 9 |
+
|
| 10 |
+
# User-friendly check for sphinx-build
|
| 11 |
+
ifeq ($(shell which $(SPHINXBUILD) >/dev/null 2>&1; echo $$?), 1)
|
| 12 |
+
$(error The '$(SPHINXBUILD)' command was not found. Make sure you have Sphinx installed, then set the SPHINXBUILD environment variable to point to the full path of the '$(SPHINXBUILD)' executable. Alternatively you can add the directory with the executable to your PATH. If you don't have Sphinx installed, grab it from http://sphinx-doc.org/)
|
| 13 |
+
endif
|
| 14 |
+
|
| 15 |
+
# Internal variables.
|
| 16 |
+
PAPEROPT_a4 = -D latex_paper_size=a4
|
| 17 |
+
PAPEROPT_letter = -D latex_paper_size=letter
|
| 18 |
+
ALLSPHINXOPTS = -d $(BUILDDIR)/doctrees $(PAPEROPT_$(PAPER)) $(SPHINXOPTS) .
|
| 19 |
+
# the i18n builder cannot share the environment and doctrees with the others
|
| 20 |
+
I18NSPHINXOPTS = $(PAPEROPT_$(PAPER)) $(SPHINXOPTS) .
|
| 21 |
+
|
| 22 |
+
.PHONY: help clean html dirhtml singlehtml pickle json htmlhelp qthelp devhelp epub latex latexpdf text man changes linkcheck doctest gettext
|
| 23 |
+
|
| 24 |
+
help:
|
| 25 |
+
@echo "Please use \`make <target>' where <target> is one of"
|
| 26 |
+
@echo " html to make standalone HTML files"
|
| 27 |
+
@echo " dirhtml to make HTML files named index.html in directories"
|
| 28 |
+
@echo " singlehtml to make a single large HTML file"
|
| 29 |
+
@echo " pickle to make pickle files"
|
| 30 |
+
@echo " json to make JSON files"
|
| 31 |
+
@echo " htmlhelp to make HTML files and a HTML help project"
|
| 32 |
+
@echo " qthelp to make HTML files and a qthelp project"
|
| 33 |
+
@echo " devhelp to make HTML files and a Devhelp project"
|
| 34 |
+
@echo " epub to make an epub"
|
| 35 |
+
@echo " latex to make LaTeX files, you can set PAPER=a4 or PAPER=letter"
|
| 36 |
+
@echo " latexpdf to make LaTeX files and run them through pdflatex"
|
| 37 |
+
@echo " latexpdfja to make LaTeX files and run them through platex/dvipdfmx"
|
| 38 |
+
@echo " text to make text files"
|
| 39 |
+
@echo " man to make manual pages"
|
| 40 |
+
@echo " texinfo to make Texinfo files"
|
| 41 |
+
@echo " info to make Texinfo files and run them through makeinfo"
|
| 42 |
+
@echo " gettext to make PO message catalogs"
|
| 43 |
+
@echo " changes to make an overview of all changed/added/deprecated items"
|
| 44 |
+
@echo " xml to make Docutils-native XML files"
|
| 45 |
+
@echo " pseudoxml to make pseudoxml-XML files for display purposes"
|
| 46 |
+
@echo " linkcheck to check all external links for integrity"
|
| 47 |
+
@echo " doctest to run all doctests embedded in the documentation (if enabled)"
|
| 48 |
+
|
| 49 |
+
clean:
|
| 50 |
+
rm -rf $(BUILDDIR)/*
|
| 51 |
+
|
| 52 |
+
html:
|
| 53 |
+
$(SPHINXBUILD) -b html $(ALLSPHINXOPTS) $(BUILDDIR)/html
|
| 54 |
+
@echo
|
| 55 |
+
@echo "Build finished. The HTML pages are in $(BUILDDIR)/html."
|
| 56 |
+
|
| 57 |
+
dirhtml:
|
| 58 |
+
$(SPHINXBUILD) -b dirhtml $(ALLSPHINXOPTS) $(BUILDDIR)/dirhtml
|
| 59 |
+
@echo
|
| 60 |
+
@echo "Build finished. The HTML pages are in $(BUILDDIR)/dirhtml."
|
| 61 |
+
|
| 62 |
+
singlehtml:
|
| 63 |
+
$(SPHINXBUILD) -b singlehtml $(ALLSPHINXOPTS) $(BUILDDIR)/singlehtml
|
| 64 |
+
@echo
|
| 65 |
+
@echo "Build finished. The HTML page is in $(BUILDDIR)/singlehtml."
|
| 66 |
+
|
| 67 |
+
pickle:
|
| 68 |
+
$(SPHINXBUILD) -b pickle $(ALLSPHINXOPTS) $(BUILDDIR)/pickle
|
| 69 |
+
@echo
|
| 70 |
+
@echo "Build finished; now you can process the pickle files."
|
| 71 |
+
|
| 72 |
+
json:
|
| 73 |
+
$(SPHINXBUILD) -b json $(ALLSPHINXOPTS) $(BUILDDIR)/json
|
| 74 |
+
@echo
|
| 75 |
+
@echo "Build finished; now you can process the JSON files."
|
| 76 |
+
|
| 77 |
+
htmlhelp:
|
| 78 |
+
$(SPHINXBUILD) -b htmlhelp $(ALLSPHINXOPTS) $(BUILDDIR)/htmlhelp
|
| 79 |
+
@echo
|
| 80 |
+
@echo "Build finished; now you can run HTML Help Workshop with the" \
|
| 81 |
+
".hhp project file in $(BUILDDIR)/htmlhelp."
|
| 82 |
+
|
| 83 |
+
qthelp:
|
| 84 |
+
$(SPHINXBUILD) -b qthelp $(ALLSPHINXOPTS) $(BUILDDIR)/qthelp
|
| 85 |
+
@echo
|
| 86 |
+
@echo "Build finished; now you can run "qcollectiongenerator" with the" \
|
| 87 |
+
".qhcp project file in $(BUILDDIR)/qthelp, like this:"
|
| 88 |
+
@echo "# qcollectiongenerator $(BUILDDIR)/qthelp/HodginHuxleyLEMSTutorial.qhcp"
|
| 89 |
+
@echo "To view the help file:"
|
| 90 |
+
@echo "# assistant -collectionFile $(BUILDDIR)/qthelp/HodginHuxleyLEMSTutorial.qhc"
|
| 91 |
+
|
| 92 |
+
devhelp:
|
| 93 |
+
$(SPHINXBUILD) -b devhelp $(ALLSPHINXOPTS) $(BUILDDIR)/devhelp
|
| 94 |
+
@echo
|
| 95 |
+
@echo "Build finished."
|
| 96 |
+
@echo "To view the help file:"
|
| 97 |
+
@echo "# mkdir -p $$HOME/.local/share/devhelp/HodginHuxleyLEMSTutorial"
|
| 98 |
+
@echo "# ln -s $(BUILDDIR)/devhelp $$HOME/.local/share/devhelp/HodginHuxleyLEMSTutorial"
|
| 99 |
+
@echo "# devhelp"
|
| 100 |
+
|
| 101 |
+
epub:
|
| 102 |
+
$(SPHINXBUILD) -b epub $(ALLSPHINXOPTS) $(BUILDDIR)/epub
|
| 103 |
+
@echo
|
| 104 |
+
@echo "Build finished. The epub file is in $(BUILDDIR)/epub."
|
| 105 |
+
|
| 106 |
+
latex:
|
| 107 |
+
$(SPHINXBUILD) -b latex $(ALLSPHINXOPTS) $(BUILDDIR)/latex
|
| 108 |
+
@echo
|
| 109 |
+
@echo "Build finished; the LaTeX files are in $(BUILDDIR)/latex."
|
| 110 |
+
@echo "Run \`make' in that directory to run these through (pdf)latex" \
|
| 111 |
+
"(use \`make latexpdf' here to do that automatically)."
|
| 112 |
+
|
| 113 |
+
latexpdf:
|
| 114 |
+
$(SPHINXBUILD) -b latex $(ALLSPHINXOPTS) $(BUILDDIR)/latex
|
| 115 |
+
@echo "Running LaTeX files through pdflatex..."
|
| 116 |
+
$(MAKE) -C $(BUILDDIR)/latex all-pdf
|
| 117 |
+
@echo "pdflatex finished; the PDF files are in $(BUILDDIR)/latex."
|
| 118 |
+
|
| 119 |
+
latexpdfja:
|
| 120 |
+
$(SPHINXBUILD) -b latex $(ALLSPHINXOPTS) $(BUILDDIR)/latex
|
| 121 |
+
@echo "Running LaTeX files through platex and dvipdfmx..."
|
| 122 |
+
$(MAKE) -C $(BUILDDIR)/latex all-pdf-ja
|
| 123 |
+
@echo "pdflatex finished; the PDF files are in $(BUILDDIR)/latex."
|
| 124 |
+
|
| 125 |
+
text:
|
| 126 |
+
$(SPHINXBUILD) -b text $(ALLSPHINXOPTS) $(BUILDDIR)/text
|
| 127 |
+
@echo
|
| 128 |
+
@echo "Build finished. The text files are in $(BUILDDIR)/text."
|
| 129 |
+
|
| 130 |
+
man:
|
| 131 |
+
$(SPHINXBUILD) -b man $(ALLSPHINXOPTS) $(BUILDDIR)/man
|
| 132 |
+
@echo
|
| 133 |
+
@echo "Build finished. The manual pages are in $(BUILDDIR)/man."
|
| 134 |
+
|
| 135 |
+
texinfo:
|
| 136 |
+
$(SPHINXBUILD) -b texinfo $(ALLSPHINXOPTS) $(BUILDDIR)/texinfo
|
| 137 |
+
@echo
|
| 138 |
+
@echo "Build finished. The Texinfo files are in $(BUILDDIR)/texinfo."
|
| 139 |
+
@echo "Run \`make' in that directory to run these through makeinfo" \
|
| 140 |
+
"(use \`make info' here to do that automatically)."
|
| 141 |
+
|
| 142 |
+
info:
|
| 143 |
+
$(SPHINXBUILD) -b texinfo $(ALLSPHINXOPTS) $(BUILDDIR)/texinfo
|
| 144 |
+
@echo "Running Texinfo files through makeinfo..."
|
| 145 |
+
make -C $(BUILDDIR)/texinfo info
|
| 146 |
+
@echo "makeinfo finished; the Info files are in $(BUILDDIR)/texinfo."
|
| 147 |
+
|
| 148 |
+
gettext:
|
| 149 |
+
$(SPHINXBUILD) -b gettext $(I18NSPHINXOPTS) $(BUILDDIR)/locale
|
| 150 |
+
@echo
|
| 151 |
+
@echo "Build finished. The message catalogs are in $(BUILDDIR)/locale."
|
| 152 |
+
|
| 153 |
+
changes:
|
| 154 |
+
$(SPHINXBUILD) -b changes $(ALLSPHINXOPTS) $(BUILDDIR)/changes
|
| 155 |
+
@echo
|
| 156 |
+
@echo "The overview file is in $(BUILDDIR)/changes."
|
| 157 |
+
|
| 158 |
+
linkcheck:
|
| 159 |
+
$(SPHINXBUILD) -b linkcheck $(ALLSPHINXOPTS) $(BUILDDIR)/linkcheck
|
| 160 |
+
@echo
|
| 161 |
+
@echo "Link check complete; look for any errors in the above output " \
|
| 162 |
+
"or in $(BUILDDIR)/linkcheck/output.txt."
|
| 163 |
+
|
| 164 |
+
doctest:
|
| 165 |
+
$(SPHINXBUILD) -b doctest $(ALLSPHINXOPTS) $(BUILDDIR)/doctest
|
| 166 |
+
@echo "Testing of doctests in the sources finished, look at the " \
|
| 167 |
+
"results in $(BUILDDIR)/doctest/output.txt."
|
| 168 |
+
|
| 169 |
+
xml:
|
| 170 |
+
$(SPHINXBUILD) -b xml $(ALLSPHINXOPTS) $(BUILDDIR)/xml
|
| 171 |
+
@echo
|
| 172 |
+
@echo "Build finished. The XML files are in $(BUILDDIR)/xml."
|
| 173 |
+
|
| 174 |
+
pseudoxml:
|
| 175 |
+
$(SPHINXBUILD) -b pseudoxml $(ALLSPHINXOPTS) $(BUILDDIR)/pseudoxml
|
| 176 |
+
@echo
|
| 177 |
+
@echo "Build finished. The pseudo-XML files are in $(BUILDDIR)/pseudoxml."
|
Tutorial/Source/GEPPETTO.md
ADDED
|
@@ -0,0 +1,12 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
Some commands to help execute this model on GEPPETTO (under OpenSourceBrain)
|
| 2 |
+
|
| 3 |
+
**Still being tested!**
|
| 4 |
+
|
| 5 |
+
HHCellNetwork.electrical.getSimulationTree();
|
| 6 |
+
|
| 7 |
+
Project.persist();
|
| 8 |
+
|
| 9 |
+
HHCellNetwork.electrical.SimulationTree.hhpop[0].v.setWatched(true);
|
| 10 |
+
|
| 11 |
+
- Press Run
|
| 12 |
+
|
Tutorial/Source/GEPPETTO.ses.js
ADDED
|
@@ -0,0 +1,35 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
/*****
|
| 2 |
+
|
| 3 |
+
A file to create info widgets, plots, save variables etc. in OSB
|
| 4 |
+
|
| 5 |
+
*****/
|
| 6 |
+
|
| 7 |
+
G.addWidget(Widgets.POPUP);
|
| 8 |
+
Popup1.setMessage("The <b>Hodgkin-Huxley model</b> is a mathematical model that describes how action potentials in neurons are initiated and propagated. It is a set of nonlinear differential equations that approximates the electrical characteristics of excitable cells such as neurons. <br/><br/>You can run your own simulations of this model by signing up to OSB and logging in. <br/><br/>There is also <a target='_blank' href='http://hodgkin-huxley-tutorial.readthedocs.io/en/latest/'>a tutorial for the HH model</a>, which has been developed as part of the <a target='_blank' href='http://www.openworm.org/'>OpenWorm project</a>.");
|
| 9 |
+
Popup1.setName("Description");
|
| 10 |
+
Popup1.setPosition(1074,142)
|
| 11 |
+
Popup1.setSize(391.8,454.8)
|
| 12 |
+
|
| 13 |
+
|
| 14 |
+
var Plot1 = G.addWidget(Widgets.PLOT);
|
| 15 |
+
Plot1.setName("Hodgkin-Huxley Spiking Neuron");
|
| 16 |
+
|
| 17 |
+
Plot1.setPosition(120, 90);
|
| 18 |
+
Plot1.setSize(230,465);
|
| 19 |
+
Plot1.plotData(HHCellNetwork.hhpop[0].v);
|
| 20 |
+
|
| 21 |
+
var Plot2 = G.addWidget(Widgets.PLOT);
|
| 22 |
+
|
| 23 |
+
Plot2.setName("Gating Variables");
|
| 24 |
+
Plot2.setPosition(120,350);
|
| 25 |
+
Plot2.setSize(285,465)
|
| 26 |
+
Plot2.plotData(HHCellNetwork.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.h.q);
|
| 27 |
+
Plot2.plotData(HHCellNetwork.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.m.q);
|
| 28 |
+
Plot2.plotData(HHCellNetwork.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.n.q);
|
| 29 |
+
|
| 30 |
+
Plot2.setLegend(HHCellNetwork.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.h.q,"Sodium h.q");
|
| 31 |
+
Plot2.setLegend(HHCellNetwork.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.m.q,"Sodium m.q");
|
| 32 |
+
Plot2.setLegend(HHCellNetwork.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.n.q,"Potassium n.q");
|
| 33 |
+
|
| 34 |
+
HHCellNetwork.hhpop[0].v.setWatched(true);
|
| 35 |
+
HHCellNetwork.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.m.q.setWatched(true);
|
Tutorial/Source/GEPPETTO.singleap.ses.js
ADDED
|
@@ -0,0 +1,43 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
/*****
|
| 2 |
+
|
| 3 |
+
A file to create info widgets, plots, save variables etc. in OSB
|
| 4 |
+
|
| 5 |
+
*****/
|
| 6 |
+
|
| 7 |
+
G.addWidget(Widgets.POPUP);
|
| 8 |
+
Popup1.setMessage("The <b>Hodgkin-Huxley model</b> is a mathematical model that describes how action potentials in neurons are initiated and propagated. It is a set of nonlinear differential equations that approximates the electrical characteristics of excitable cells such as neurons. <br/><br/>You can run your own simulations of this model by signing up to OSB and logging in. <br/><br/>There is also <a target='_blank' href='http://hodgkin-huxley-tutorial.readthedocs.io/en/latest/'>a tutorial for the HH model</a>, which has been developed as part of the <a target='_blank' href='http://www.openworm.org/'>OpenWorm project</a>.");
|
| 9 |
+
Popup1.setName("Description");
|
| 10 |
+
Popup1.setPosition(1074,142)
|
| 11 |
+
Popup1.setSize(391.8,454.8)
|
| 12 |
+
|
| 13 |
+
|
| 14 |
+
var Plot1 = G.addWidget(Widgets.PLOT);
|
| 15 |
+
Plot1.setName("Hodgkin-Huxley Spiking Neuron");
|
| 16 |
+
|
| 17 |
+
Plot1.setPosition(120, 90);
|
| 18 |
+
Plot1.setSize(230,465);
|
| 19 |
+
Plot1.plotData(HHCellSingleAP.hhpop[0].v);
|
| 20 |
+
|
| 21 |
+
var Plot2 = G.addWidget(Widgets.PLOT);
|
| 22 |
+
|
| 23 |
+
Plot2.setName("Gating Variables");
|
| 24 |
+
Plot2.setPosition(120,350);
|
| 25 |
+
Plot2.setSize(285,465)
|
| 26 |
+
Plot2.plotData(HHCellSingleAP.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.h.q);
|
| 27 |
+
Plot2.plotData(HHCellSingleAP.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.m.q);
|
| 28 |
+
Plot2.plotData(HHCellSingleAP.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.n.q);
|
| 29 |
+
|
| 30 |
+
Plot2.setLegend(HHCellSingleAP.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.h.q,"Sodium h.q");
|
| 31 |
+
Plot2.setLegend(HHCellSingleAP.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.m.q,"Sodium m.q");
|
| 32 |
+
Plot2.setLegend(HHCellSingleAP.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.n.q,"Potassium n.q");
|
| 33 |
+
|
| 34 |
+
|
| 35 |
+
Instances.getInstance("HHCellSingleAP.hhpop[0].v");
|
| 36 |
+
HHCellSingleAP.hhpop[0].v.setWatched(true);
|
| 37 |
+
|
| 38 |
+
Instances.getInstance("HHCellSingleAP.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.m.q");
|
| 39 |
+
HHCellSingleAP.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.m.q.setWatched(true);
|
| 40 |
+
Instances.getInstance("HHCellSingleAP.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.h.q");
|
| 41 |
+
HHCellSingleAP.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.h.q.setWatched(true);
|
| 42 |
+
Instances.getInstance("HHCellSingleAP.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.n.q");
|
| 43 |
+
HHCellSingleAP.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.n.q.setWatched(true);
|
Tutorial/Source/GEPPETTO.vclamp.ses.js
ADDED
|
@@ -0,0 +1,59 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
/*****
|
| 2 |
+
|
| 3 |
+
A file to create info widgets, plots, save variables etc. in OSB
|
| 4 |
+
|
| 5 |
+
*****/
|
| 6 |
+
|
| 7 |
+
G.addWidget(Widgets.POPUP);
|
| 8 |
+
Popup1.setMessage("The <b>Hodgkin-Huxley model</b> is a mathematical model that describes how action potentials in neurons are initiated and propagated. It is a set of nonlinear differential equations that approximates the electrical characteristics of excitable cells such as neurons. <br/><br/>You can run your own simulations of this model by signing up to OSB and logging in. <br/><br/>There is also <a target='_blank' href='http://hodgkin-huxley-tutorial.readthedocs.io/en/latest/'>a tutorial for the HH model</a>, which has been developed as part of the <a target='_blank' href='http://www.openworm.org/'>OpenWorm project</a>.");
|
| 9 |
+
Popup1.setName("Description");
|
| 10 |
+
Popup1.setPosition(1074,142)
|
| 11 |
+
Popup1.setSize(391.8,454.8)
|
| 12 |
+
|
| 13 |
+
|
| 14 |
+
var Plot1 = G.addWidget(Widgets.PLOT);
|
| 15 |
+
Plot1.setName("Hodgkin-Huxley Spiking Neuron");
|
| 16 |
+
|
| 17 |
+
Plot1.setPosition(120, 90);
|
| 18 |
+
Plot1.setSize(230,465);
|
| 19 |
+
Plot1.plotData(HHCellVClamp.hhpop[0].v);
|
| 20 |
+
|
| 21 |
+
var Plot2 = G.addWidget(Widgets.PLOT);
|
| 22 |
+
|
| 23 |
+
Plot2.setName("Gating Variables");
|
| 24 |
+
Plot2.setPosition(120,350);
|
| 25 |
+
Plot2.setSize(285,465)
|
| 26 |
+
Plot2.plotData(HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.h.q);
|
| 27 |
+
Plot2.plotData(HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.m.q);
|
| 28 |
+
Plot2.plotData(HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.n.q);
|
| 29 |
+
|
| 30 |
+
Plot2.setLegend(HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.h.q,"Sodium h.q");
|
| 31 |
+
Plot2.setLegend(HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.m.q,"Sodium m.q");
|
| 32 |
+
Plot2.setLegend(HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.n.q,"Potassium n.q");
|
| 33 |
+
|
| 34 |
+
var Plot3 = G.addWidget(Widgets.PLOT);
|
| 35 |
+
|
| 36 |
+
Plot3.setName("Conductances");
|
| 37 |
+
Plot3.setPosition(120,350);
|
| 38 |
+
Plot3.setSize(285,465)
|
| 39 |
+
Plot3.plotData(HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.g);
|
| 40 |
+
Plot3.setLegend(HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.g,"Sodium g");
|
| 41 |
+
Plot3.plotData(HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.g);
|
| 42 |
+
Plot3.setLegend(HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.g,"Potassium g");
|
| 43 |
+
|
| 44 |
+
|
| 45 |
+
|
| 46 |
+
Instances.getInstance("HHCellVClamp.hhpop[0].v");
|
| 47 |
+
HHCellVClamp.hhpop[0].v.setWatched(true);
|
| 48 |
+
|
| 49 |
+
Instances.getInstance("HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.m.q");
|
| 50 |
+
HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.m.q.setWatched(true);
|
| 51 |
+
Instances.getInstance("HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.h.q");
|
| 52 |
+
HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.h.q.setWatched(true);
|
| 53 |
+
Instances.getInstance("HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.n.q");
|
| 54 |
+
HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.n.q.setWatched(true);
|
| 55 |
+
|
| 56 |
+
Instances.getInstance("HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.g");
|
| 57 |
+
HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.naChans.naChan.g.setWatched(true);
|
| 58 |
+
Instances.getInstance("HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.g");
|
| 59 |
+
HHCellVClamp.hhpop[0].bioPhys1.membraneProperties.kChans.kChan.g.setWatched(true);
|
Tutorial/Source/HHCellNetwork.net.nml
ADDED
|
@@ -0,0 +1,24 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<?xml version="1.0" encoding="UTF-8"?>
|
| 2 |
+
|
| 3 |
+
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2"
|
| 4 |
+
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
|
| 5 |
+
xsi:schemaLocation="http://www.neuroml.org/schema/neuroml2 https://raw.githubusercontent.com/NeuroML/NeuroML2/master/Schemas/NeuroML2/NeuroML_v2beta3.xsd"
|
| 6 |
+
id="HHCellNetwork">
|
| 7 |
+
|
| 8 |
+
<include href="hhcell.cell.nml"/> <!-- Include the cell definition -->
|
| 9 |
+
|
| 10 |
+
<!-- Short small current pulse input & short larger current pulse input -->
|
| 11 |
+
<pulseGenerator id="pulseGen1" delay="100ms" duration="100ms" amplitude="0.10nA"/>
|
| 12 |
+
<pulseGenerator id="pulseGen2" delay="300ms" duration="100ms" amplitude="0.35nA"/>
|
| 13 |
+
|
| 14 |
+
<network id="HHCellNetwork">
|
| 15 |
+
|
| 16 |
+
<notes>Network with a single cell based on the Hodgkin Huxley model with 2 input currents</notes>
|
| 17 |
+
|
| 18 |
+
<population id="hhpop" component="hhcell" size="1"/>
|
| 19 |
+
<explicitInput target="hhpop[0]" input="pulseGen1"/>
|
| 20 |
+
<explicitInput target="hhpop[0]" input="pulseGen2"/>
|
| 21 |
+
</network>
|
| 22 |
+
|
| 23 |
+
</neuroml>
|
| 24 |
+
|
Tutorial/Source/HHCellSingleAP.net.nml
ADDED
|
@@ -0,0 +1,36 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<?xml version="1.0" encoding="UTF-8"?>
|
| 2 |
+
|
| 3 |
+
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2"
|
| 4 |
+
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
|
| 5 |
+
xsi:schemaLocation="http://www.neuroml.org/schema/neuroml2 https://raw.githubusercontent.com/NeuroML/NeuroML2/master/Schemas/NeuroML2/NeuroML_v2beta3.xsd"
|
| 6 |
+
id="HHCellNetwork">
|
| 7 |
+
|
| 8 |
+
<include href="hhcell.cell.nml"/> <!-- Include the cell definition -->
|
| 9 |
+
|
| 10 |
+
<!-- Short small current pulse input & short larger current pulse input -->
|
| 11 |
+
<pulseGenerator id="pulseGen1" delay="5ms" duration="25ms" amplitude="0.05nA"/>
|
| 12 |
+
|
| 13 |
+
<network id="HHCellNetwork">
|
| 14 |
+
|
| 15 |
+
<notes>Network with a single cell based on the Hodgkin Huxley model with 1 input current</notes>
|
| 16 |
+
|
| 17 |
+
<property tag="recommended_dt_ms" value="0.025"/>
|
| 18 |
+
<property tag="recommended_duration_ms" value="50.0"/>
|
| 19 |
+
|
| 20 |
+
<population id="hhpop" component="hhcell" type="populationList" size="1">
|
| 21 |
+
<annotation>
|
| 22 |
+
<property tag="color" value="1 0 0"/>
|
| 23 |
+
</annotation>
|
| 24 |
+
<instance id="0">
|
| 25 |
+
<location x="0" y="0" z="0"/>
|
| 26 |
+
</instance>
|
| 27 |
+
</population>
|
| 28 |
+
|
| 29 |
+
<inputList id="Input_0" component="pulseGen1" population="hhpop">
|
| 30 |
+
<input id="0" target="../hhpop/0/hhcell" destination="synapses"/>
|
| 31 |
+
</inputList>
|
| 32 |
+
|
| 33 |
+
</network>
|
| 34 |
+
|
| 35 |
+
</neuroml>
|
| 36 |
+
|
Tutorial/Source/HHCellVClamp.net.nml
ADDED
|
@@ -0,0 +1,23 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<?xml version="1.0" encoding="UTF-8"?>
|
| 2 |
+
|
| 3 |
+
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2"
|
| 4 |
+
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
|
| 5 |
+
xsi:schemaLocation="http://www.neuroml.org/schema/neuroml2 https://raw.githubusercontent.com/NeuroML/NeuroML2/master/Schemas/NeuroML2/NeuroML_v2beta3.xsd"
|
| 6 |
+
id="HHCellVClamp">
|
| 7 |
+
|
| 8 |
+
<include href="hhcell.cell.nml"/> <!-- Include the cell definition -->
|
| 9 |
+
<include href="vclamp.xml"/> <!-- Include the voltage clamp definition -->
|
| 10 |
+
|
| 11 |
+
<network id="HHCellNetwork">
|
| 12 |
+
|
| 13 |
+
<notes>Network with a single cell based on the Hodgkin Huxley model with a voltage clamp</notes>
|
| 14 |
+
|
| 15 |
+
<population id="hhpop" component="hhcell" size="1"/>
|
| 16 |
+
<explicitInput target="hhpop[0]" input="vClamp"/>
|
| 17 |
+
|
| 18 |
+
</network>
|
| 19 |
+
|
| 20 |
+
|
| 21 |
+
|
| 22 |
+
</neuroml>
|
| 23 |
+
|
Tutorial/Source/HodgkinHuxley.py
ADDED
|
@@ -0,0 +1,401 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
import scipy as sp
|
| 2 |
+
import numpy as np
|
| 3 |
+
import pylab as plt
|
| 4 |
+
from scipy.integrate import odeint
|
| 5 |
+
import sys
|
| 6 |
+
|
| 7 |
+
class HodgkinHuxley():
|
| 8 |
+
"""Full Hodgkin-Huxley Model implemented in Python"""
|
| 9 |
+
|
| 10 |
+
""" __init__ uses optional arguments """
|
| 11 |
+
""" when no argument is passed default values are used """
|
| 12 |
+
|
| 13 |
+
def __init__(self, C_m=1, gmax_Na=120, gmax_K=36, gmax_L=0.3, E_Na=50,
|
| 14 |
+
E_K=-77, E_L=-54.387, t_n=450, delta_t=0.01,
|
| 15 |
+
I_inj_amplitude=0, I_inj_duration=0, I_inj_delay=0,
|
| 16 |
+
vc_delay=10, vc_duration=30, vc_condVoltage=-65,
|
| 17 |
+
vc_testVoltage=10, vc_returnVoltage=-65, runMode='iclamp',
|
| 18 |
+
injected_current_plot=True, gating_plot=True, cond_scaling_plot=False,
|
| 19 |
+
cond_dens_plot=True, driving_force_plot=False,
|
| 20 |
+
current_plot=True, memb_pot_plot=True):
|
| 21 |
+
|
| 22 |
+
self.C_m = C_m
|
| 23 |
+
""" membrane capacitance, in uF/cm^2 """
|
| 24 |
+
|
| 25 |
+
self.gmax_Na = gmax_Na
|
| 26 |
+
""" Sodium (Na) maximum conductances, in mS/cm^2 """
|
| 27 |
+
|
| 28 |
+
self.gmax_K = gmax_K
|
| 29 |
+
""" Postassium (K) maximum conductances, in mS/cm^2 """
|
| 30 |
+
|
| 31 |
+
self.gmax_L = gmax_L
|
| 32 |
+
""" Leak maximum conductances, in mS/cm^2 """
|
| 33 |
+
|
| 34 |
+
self.E_Na = E_Na
|
| 35 |
+
""" Sodium (Na) Nernst reversal potentials, in mV """
|
| 36 |
+
|
| 37 |
+
self.E_K = E_K
|
| 38 |
+
""" Postassium (K) Nernst reversal potentials, in mV """
|
| 39 |
+
|
| 40 |
+
self.E_L = E_L
|
| 41 |
+
""" Leak Nernst reversal potentials, in mV """
|
| 42 |
+
|
| 43 |
+
self.t = np.arange(0, t_n, delta_t)
|
| 44 |
+
""" The time to integrate over """
|
| 45 |
+
|
| 46 |
+
""" Advanced input - injection current (single rectangular pulse only) """
|
| 47 |
+
|
| 48 |
+
self.I_inj_amplitude = I_inj_amplitude
|
| 49 |
+
""" maximum value or amplitude of injection pulse """
|
| 50 |
+
|
| 51 |
+
self.I_inj_duration = I_inj_duration
|
| 52 |
+
""" duration or width of injection pulse """
|
| 53 |
+
|
| 54 |
+
self.I_inj_delay = I_inj_delay
|
| 55 |
+
""" start time of injection pulse """
|
| 56 |
+
|
| 57 |
+
#vclamp parameters
|
| 58 |
+
self.run_mode = runMode
|
| 59 |
+
"""default is current clamp"""
|
| 60 |
+
|
| 61 |
+
self.delay = vc_delay
|
| 62 |
+
"""Delay before switching from conditioningVoltage to testingVoltage, in ms"""
|
| 63 |
+
|
| 64 |
+
self.duration = vc_duration
|
| 65 |
+
"""Duration to hold at testingVoltage, in ms"""
|
| 66 |
+
|
| 67 |
+
self.conditioningVoltage = vc_condVoltage
|
| 68 |
+
"""Target voltage before time delay, in mV"""
|
| 69 |
+
|
| 70 |
+
self.testingVoltage = vc_testVoltage
|
| 71 |
+
"""Target voltage between times delay and delay + duration, in mV"""
|
| 72 |
+
|
| 73 |
+
self.returnVoltage = vc_returnVoltage
|
| 74 |
+
"""Target voltage after time duration, in mV"""
|
| 75 |
+
|
| 76 |
+
self.simpleSeriesResistance = 1e7
|
| 77 |
+
"""Current will be calculated by the difference in voltage between the target and parent, divided by this value, in mOhm"""
|
| 78 |
+
|
| 79 |
+
# plotting conditionals
|
| 80 |
+
self.injected_current_plot = injected_current_plot
|
| 81 |
+
self.gating_plot = gating_plot
|
| 82 |
+
self.cond_scaling_plot = cond_scaling_plot
|
| 83 |
+
self.cond_dens_plot = cond_dens_plot
|
| 84 |
+
self.driving_force_plot = driving_force_plot
|
| 85 |
+
self.current_plot = current_plot
|
| 86 |
+
self.memb_pot_plot = memb_pot_plot
|
| 87 |
+
|
| 88 |
+
self.num_plots = (int(self.injected_current_plot) +
|
| 89 |
+
int(self.gating_plot)+ int(self.cond_scaling_plot) +
|
| 90 |
+
int(self.cond_dens_plot) + int(self.driving_force_plot) +
|
| 91 |
+
int(self.current_plot) + int(self.memb_pot_plot))
|
| 92 |
+
|
| 93 |
+
self.plot_count = 0
|
| 94 |
+
|
| 95 |
+
def alpha_m(self, V):
|
| 96 |
+
"""Channel gating kinetics. Functions of membrane voltage"""
|
| 97 |
+
return 0.1*(V+40.0)/(1.0 - np.exp(-(V+40.0) / 10.0))
|
| 98 |
+
|
| 99 |
+
def beta_m(self, V):
|
| 100 |
+
"""Channel gating kinetics. Functions of membrane voltage"""
|
| 101 |
+
return 4.0*np.exp(-(V+65.0) / 18.0)
|
| 102 |
+
|
| 103 |
+
def alpha_h(self, V):
|
| 104 |
+
"""Channel gating kinetics. Functions of membrane voltage"""
|
| 105 |
+
return 0.07*np.exp(-(V+65.0) / 20.0)
|
| 106 |
+
|
| 107 |
+
def beta_h(self, V):
|
| 108 |
+
"""Channel gating kinetics. Functions of membrane voltage"""
|
| 109 |
+
return 1.0/(1.0 + np.exp(-(V+35.0) / 10.0))
|
| 110 |
+
|
| 111 |
+
def alpha_n(self, V):
|
| 112 |
+
"""Channel gating kinetics. Functions of membrane voltage"""
|
| 113 |
+
return 0.01*(V+55.0)/(1.0 - np.exp(-(V+55.0) / 10.0))
|
| 114 |
+
|
| 115 |
+
def beta_n(self, V):
|
| 116 |
+
"""Channel gating kinetics. Functions of membrane voltage"""
|
| 117 |
+
return 0.125*np.exp(-(V+65) / 80.0)
|
| 118 |
+
|
| 119 |
+
def g_Na(self, m, h):
|
| 120 |
+
"""
|
| 121 |
+
Conductance density (in mS/cm^2)
|
| 122 |
+
Sodium (Na = element name)
|
| 123 |
+
|
| 124 |
+
| :param m:
|
| 125 |
+
| :param h:
|
| 126 |
+
| :return:
|
| 127 |
+
"""
|
| 128 |
+
return self.gmax_Na * m**3 * h
|
| 129 |
+
|
| 130 |
+
def I_Na(self, V, m, h):
|
| 131 |
+
"""
|
| 132 |
+
Membrane current (in uA/cm^2)
|
| 133 |
+
Sodium (Na = element name)
|
| 134 |
+
|
| 135 |
+
| :param V:
|
| 136 |
+
| :param m:
|
| 137 |
+
| :param h:
|
| 138 |
+
| :return:
|
| 139 |
+
"""
|
| 140 |
+
return self.g_Na(m, h) * (V - self.E_Na)
|
| 141 |
+
|
| 142 |
+
|
| 143 |
+
def g_K(self, n):
|
| 144 |
+
"""
|
| 145 |
+
Conductance density (in mS/cm^2)
|
| 146 |
+
Potassium (K = element name)
|
| 147 |
+
|
| 148 |
+
| :param n:
|
| 149 |
+
| :return:
|
| 150 |
+
"""
|
| 151 |
+
return self.gmax_K * n**4
|
| 152 |
+
|
| 153 |
+
def I_K(self, V, n):
|
| 154 |
+
"""
|
| 155 |
+
Membrane current (in uA/cm^2)
|
| 156 |
+
Potassium (K = element name)
|
| 157 |
+
|
| 158 |
+
| :param V:
|
| 159 |
+
| :param n:
|
| 160 |
+
| :return:
|
| 161 |
+
"""
|
| 162 |
+
return self.g_K(n) * (V - self.E_K)
|
| 163 |
+
|
| 164 |
+
# Leak
|
| 165 |
+
def I_L(self, V):
|
| 166 |
+
"""
|
| 167 |
+
Membrane current (in uA/cm^2)
|
| 168 |
+
Leak
|
| 169 |
+
|
| 170 |
+
| :param V:
|
| 171 |
+
| :param h:
|
| 172 |
+
| :return:
|
| 173 |
+
"""
|
| 174 |
+
return self.gmax_L * (V - self.E_L)
|
| 175 |
+
|
| 176 |
+
def I_inj(self, t):
|
| 177 |
+
"""
|
| 178 |
+
External Current
|
| 179 |
+
|
| 180 |
+
| :param t: time
|
| 181 |
+
| :return: step up to 10 uA/cm^2 at t>100
|
| 182 |
+
| step down to 0 uA/cm^2 at t>200
|
| 183 |
+
| step up to 35 uA/cm^2 at t>300
|
| 184 |
+
| step down to 0 uA/cm^2 at t>400
|
| 185 |
+
"""
|
| 186 |
+
|
| 187 |
+
""" running standalone python script """
|
| 188 |
+
if __name__ == '__main__':
|
| 189 |
+
return 10*(t>100) - 10*(t>200) + 35*(t>300) - 35*(t>400)
|
| 190 |
+
|
| 191 |
+
#""" running jupyterLab notebook """
|
| 192 |
+
else:
|
| 193 |
+
return self.I_inj_amplitude*(t>self.I_inj_delay) - self.I_inj_amplitude*(t>self.I_inj_delay+self.I_inj_duration)
|
| 194 |
+
|
| 195 |
+
def I_inj_vclamp(self,t,v):
|
| 196 |
+
"""
|
| 197 |
+
External Current (vclamp)
|
| 198 |
+
|
| 199 |
+
| :param t: time
|
| 200 |
+
| :return: injector current for voltage clamp
|
| 201 |
+
|
|
| 202 |
+
"""
|
| 203 |
+
if t > (self.delay + self.duration):
|
| 204 |
+
current_A = (self.returnVoltage - v) / self.simpleSeriesResistance
|
| 205 |
+
elif t >= self.delay:
|
| 206 |
+
current_A = (self.testingVoltage - v) / self.simpleSeriesResistance
|
| 207 |
+
elif t < self.delay:
|
| 208 |
+
current_A = (self.conditioningVoltage - v) / self.simpleSeriesResistance
|
| 209 |
+
else:
|
| 210 |
+
print('Problem in injection current calculation for voltage clamp...')
|
| 211 |
+
return 0
|
| 212 |
+
|
| 213 |
+
#convert current to current density (uA/cm^2)
|
| 214 |
+
current_uA = current_A*10**6 #convert ampere to micro ampere
|
| 215 |
+
surface_area = 1000*10**-8 #surface area of 1000 um^2 converted to cm^2
|
| 216 |
+
current_density = current_uA/surface_area
|
| 217 |
+
|
| 218 |
+
return current_density
|
| 219 |
+
|
| 220 |
+
@staticmethod
|
| 221 |
+
def dALLdt(X, t, self):
|
| 222 |
+
"""
|
| 223 |
+
Integrate
|
| 224 |
+
|
| 225 |
+
| :param X:
|
| 226 |
+
| :param t:
|
| 227 |
+
| :return: calculate membrane potential & activation variables
|
| 228 |
+
"""
|
| 229 |
+
V, m, h, n = X
|
| 230 |
+
if self.is_vclamp():
|
| 231 |
+
dVdt = (self.I_inj_vclamp(t,V) - self.I_Na(V, m, h) - self.I_K(V, n) - self.I_L(V)) / self.C_m
|
| 232 |
+
else:
|
| 233 |
+
dVdt = (self.I_inj(t) - self.I_Na(V, m, h) - self.I_K(V, n) - self.I_L(V)) / self.C_m
|
| 234 |
+
|
| 235 |
+
dmdt = self.alpha_m(V)*(1.0-m) - self.beta_m(V)*m
|
| 236 |
+
dhdt = self.alpha_h(V)*(1.0-h) - self.beta_h(V)*h
|
| 237 |
+
dndt = self.alpha_n(V)*(1.0-n) - self.beta_n(V)*n
|
| 238 |
+
return dVdt, dmdt, dhdt, dndt
|
| 239 |
+
|
| 240 |
+
def is_vclamp(self):
|
| 241 |
+
return self.run_mode=='vclamp' or self.run_mode=='Voltage Clamp'
|
| 242 |
+
|
| 243 |
+
def simulate(self, init_values=[-64.99584, 0.05296, 0.59590, 0.31773]):
|
| 244 |
+
"""
|
| 245 |
+
Main simulate method for the Hodgkin Huxley neuron model
|
| 246 |
+
"""
|
| 247 |
+
|
| 248 |
+
# init_values are the steady state values for v,m,h,n at zero current injection
|
| 249 |
+
X = odeint(self.dALLdt, init_values, self.t, args=(self,))
|
| 250 |
+
V = X[:,0]
|
| 251 |
+
m = X[:,1]
|
| 252 |
+
h = X[:,2]
|
| 253 |
+
n = X[:,3]
|
| 254 |
+
ina = self.I_Na(V, m, h)
|
| 255 |
+
ik = self.I_K(V, n)
|
| 256 |
+
il = self.I_L(V)
|
| 257 |
+
gna = self.g_Na(m, h)
|
| 258 |
+
gk = self.g_K(n)
|
| 259 |
+
|
| 260 |
+
# Save some of the data to file
|
| 261 |
+
with open('hh_py_v.dat','w') as f:
|
| 262 |
+
for ti in range(len(self.t)):
|
| 263 |
+
f.write('%s\t%s\n'%(self.t[ti],V[ti]))
|
| 264 |
+
|
| 265 |
+
if not '-nogui' in sys.argv:
|
| 266 |
+
#increase figure and font size for display in jupyter notebook
|
| 267 |
+
|
| 268 |
+
|
| 269 |
+
if __name__ != '__main__':
|
| 270 |
+
plt.rcParams['figure.figsize'] = [7, 7]
|
| 271 |
+
#plt.rcParams['font.size'] = 15
|
| 272 |
+
#plt.rcParams['legend.fontsize'] = 12
|
| 273 |
+
plt.rcParams['legend.loc'] = "upper right"
|
| 274 |
+
#
|
| 275 |
+
else:
|
| 276 |
+
plt.rcParams['figure.figsize'] = [10, 7]
|
| 277 |
+
|
| 278 |
+
plt.close()
|
| 279 |
+
|
| 280 |
+
fig=plt.figure(figsize=(7, self.num_plots * 2))
|
| 281 |
+
fig.canvas.header_visible = False
|
| 282 |
+
# plt.xlim([np.min(self.t),np.max(self.t)]) #for all subplots
|
| 283 |
+
|
| 284 |
+
if self.injected_current_plot:
|
| 285 |
+
ax1 = plt.subplot(self.num_plots,1,self.plot_count + 1)
|
| 286 |
+
plt.title('Simulation of Hodgkin Huxley model neuron')
|
| 287 |
+
if self.is_vclamp():
|
| 288 |
+
i_inj_values = [self.I_inj_vclamp(t,v) for t,v in zip(self.t,V)]
|
| 289 |
+
else:
|
| 290 |
+
i_inj_values = [self.I_inj(t) for t in self.t]
|
| 291 |
+
|
| 292 |
+
if self.is_vclamp(): plt.ylim(-2000,3000)
|
| 293 |
+
|
| 294 |
+
plt.plot(self.t, i_inj_values, 'k')
|
| 295 |
+
plt.ylabel('$I_{inj}$ ($\\mu{A}/cm^2$)')
|
| 296 |
+
|
| 297 |
+
self.plot_count += 1
|
| 298 |
+
|
| 299 |
+
|
| 300 |
+
if self.gating_plot:
|
| 301 |
+
try:
|
| 302 |
+
plt.subplot(self.num_plots,1,self.plot_count+1, sharex = ax1)
|
| 303 |
+
except NameError:
|
| 304 |
+
ax1 = plt.subplot(self.num_plots,1,self.plot_count + 1)
|
| 305 |
+
plt.title('Simulation of Hodgkin Huxley model neuron')
|
| 306 |
+
plt.plot(self.t, m, 'r', label='$m$')
|
| 307 |
+
plt.plot(self.t, h, 'g', label='$h$')
|
| 308 |
+
plt.plot(self.t, n, 'b', label='$n$')
|
| 309 |
+
plt.ylabel('Gating variable')
|
| 310 |
+
plt.legend()
|
| 311 |
+
self.plot_count += 1
|
| 312 |
+
|
| 313 |
+
if self.cond_scaling_plot:
|
| 314 |
+
try:
|
| 315 |
+
plt.subplot(self.num_plots,1,self.plot_count+1, sharex = ax1)
|
| 316 |
+
except NameError:
|
| 317 |
+
ax1 = plt.subplot(self.num_plots,1,self.plot_count + 1)
|
| 318 |
+
plt.title('Simulation of Hodgkin Huxley model neuron')
|
| 319 |
+
scale_na = m*m*m*h
|
| 320 |
+
scale_k = n*n*n*n
|
| 321 |
+
plt.plot(self.t, scale_na, 'c', label='$m^{3}h$')
|
| 322 |
+
plt.plot(self.t, scale_k, 'y', label='$n^{4}$')
|
| 323 |
+
plt.ylabel('Cond scaling')
|
| 324 |
+
plt.legend()
|
| 325 |
+
self.plot_count += 1
|
| 326 |
+
|
| 327 |
+
if self.cond_dens_plot:
|
| 328 |
+
try:
|
| 329 |
+
plt.subplot(self.num_plots,1,self.plot_count+1, sharex = ax1)
|
| 330 |
+
except NameError:
|
| 331 |
+
ax1 = plt.subplot(self.num_plots,1,self.plot_count + 1)
|
| 332 |
+
plt.title('Simulation of Hodgkin Huxley model neuron')
|
| 333 |
+
plt.plot(self.t, gna, 'c', label='$g_{Na}$')
|
| 334 |
+
plt.plot(self.t, gk, 'y', label='$g_{K}$')
|
| 335 |
+
plt.ylabel('Cond dens ($mS/cm^2$)')
|
| 336 |
+
plt.legend()
|
| 337 |
+
self.plot_count += 1
|
| 338 |
+
|
| 339 |
+
|
| 340 |
+
if self.driving_force_plot:
|
| 341 |
+
try:
|
| 342 |
+
ax_here = plt.subplot(self.num_plots,1,self.plot_count+1, sharex = ax1)
|
| 343 |
+
except NameError:
|
| 344 |
+
ax1 = plt.subplot(self.num_plots,1,self.plot_count + 1)
|
| 345 |
+
plt.title('Simulation of Hodgkin Huxley model neuron')
|
| 346 |
+
ax_here = ax1
|
| 347 |
+
|
| 348 |
+
dna = V - self.E_Na
|
| 349 |
+
dk = V - self.E_K
|
| 350 |
+
zero = [0 for v in V]
|
| 351 |
+
|
| 352 |
+
#plt.plot(self.t, dna, 'c', label='$V - E_{Na}$')
|
| 353 |
+
ax_here.fill_between(self.t, dna, color='c', alpha=0.5)
|
| 354 |
+
ax_here.fill_between(self.t, dk, color='y', alpha=0.5)
|
| 355 |
+
|
| 356 |
+
plt.plot(self.t, dna, 'c', label='$V_{m} - E_{Na}$', linewidth=0.8)
|
| 357 |
+
plt.plot(self.t, dk, 'y', label='$V_{m} - E_{K}$', linewidth=0.8)
|
| 358 |
+
plt.plot(self.t, zero, 'k', linestyle='dashed', linewidth=0.5)
|
| 359 |
+
plt.ylabel('Driving force (mV)')
|
| 360 |
+
plt.legend()
|
| 361 |
+
#if not self.is_vclamp(): plt.ylim(-85,60)
|
| 362 |
+
#plt.ylim(-1, 40)
|
| 363 |
+
self.plot_count += 1
|
| 364 |
+
|
| 365 |
+
if self.current_plot:
|
| 366 |
+
try:
|
| 367 |
+
plt.subplot(self.num_plots,1,self.plot_count+1, sharex = ax1)
|
| 368 |
+
except NameError:
|
| 369 |
+
ax1 = plt.subplot(self.num_plots,1,self.plot_count + 1)
|
| 370 |
+
plt.title('Simulation of Hodgkin Huxley model neuron')
|
| 371 |
+
plt.plot(self.t, ina, 'c', label='$I_{Na}$')
|
| 372 |
+
plt.plot(self.t, ik, 'y', label='$I_{K}$')
|
| 373 |
+
plt.plot(self.t, il, 'm', label='$I_{L}$')
|
| 374 |
+
plt.ylabel('Curr dens ($\\mu{A}/cm^2$)')
|
| 375 |
+
plt.legend()
|
| 376 |
+
self.plot_count += 1
|
| 377 |
+
|
| 378 |
+
if self.memb_pot_plot:
|
| 379 |
+
try:
|
| 380 |
+
plt.subplot(self.num_plots,1,self.plot_count+1, sharex = ax1)
|
| 381 |
+
except NameError:
|
| 382 |
+
ax1 = plt.subplot(self.num_plots,1,self.plot_count + 1)
|
| 383 |
+
plt.title('Simulation of Hodgkin Huxley model neuron')
|
| 384 |
+
plt.plot(self.t, V, 'k')
|
| 385 |
+
plt.ylabel('$V_{m}$ (mV)')
|
| 386 |
+
plt.xlabel('Time (ms)')
|
| 387 |
+
if not self.is_vclamp(): plt.ylim(-85,60)
|
| 388 |
+
#plt.ylim(-1, 40)
|
| 389 |
+
self.plot_count += 1
|
| 390 |
+
|
| 391 |
+
plt.tight_layout()
|
| 392 |
+
plt.show()
|
| 393 |
+
|
| 394 |
+
if __name__ == '__main__':
|
| 395 |
+
|
| 396 |
+
if '-vclamp' in sys.argv:
|
| 397 |
+
runner = HodgkinHuxley(runMode='vclamp', t_n=50, delta_t=0.0005)
|
| 398 |
+
else: #default mode
|
| 399 |
+
runner = HodgkinHuxley(runMode='iclamp', t_n=450, delta_t=0.01)
|
| 400 |
+
|
| 401 |
+
runner.simulate()
|
Tutorial/Source/LEMS_HH_Simulation.xml
ADDED
|
@@ -0,0 +1,69 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<Lems>
|
| 2 |
+
|
| 3 |
+
<!-- Example with Simple Hodgkin-Huxley cell specifying segment details-->
|
| 4 |
+
|
| 5 |
+
<!-- This is a file which can be read and executed by the LEMS Interpreter.
|
| 6 |
+
It imports the LEMS definitions of the core NeuroML 2 Components,
|
| 7 |
+
imports in "pure" NeuroML 2 and contains some LEMS elements for running
|
| 8 |
+
a simulation -->
|
| 9 |
+
|
| 10 |
+
|
| 11 |
+
<Target component="sim1"/>
|
| 12 |
+
|
| 13 |
+
<Include file="Cells.xml"/>
|
| 14 |
+
<Include file="Networks.xml"/>
|
| 15 |
+
<Include file="Simulation.xml"/>
|
| 16 |
+
|
| 17 |
+
|
| 18 |
+
<Include file="HHCellNetwork.net.nml"/>
|
| 19 |
+
<Include file="hhcell.cell.nml"/>
|
| 20 |
+
<Include file="passiveChan.channel.nml"/>
|
| 21 |
+
<Include file="naChan.channel.nml"/>
|
| 22 |
+
<Include file="kChan.channel.nml"/>
|
| 23 |
+
|
| 24 |
+
|
| 25 |
+
<Simulation id="sim1" length="450ms" step="0.01ms" target="HHCellNetwork">
|
| 26 |
+
|
| 27 |
+
<Display id="d1" title="Hodgkin-Huxley Neuron: V (mV)" timeScale="1ms" xmin="-20" xmax="470" ymin="-90" ymax="50">
|
| 28 |
+
<Line id="V" quantity="hhpop[0]/v" scale="1mV" color="#000000" timeScale="1ms"/>
|
| 29 |
+
</Display>
|
| 30 |
+
|
| 31 |
+
<Display id="d2" title="Hodgkin-Huxley Neuron: Gating Variables" timeScale="1ms" xmin="-20" xmax="470" ymin="-0.1" ymax="1.1">
|
| 32 |
+
<Line id="m" quantity="hhpop[0]/bioPhys1/membraneProperties/naChans/naChan/m/q" scale="1" color="#ff0000" timeScale="1ms"/>
|
| 33 |
+
<Line id="h" quantity="hhpop[0]/bioPhys1/membraneProperties/naChans/naChan/h/q" scale="1" color="#00dd00" timeScale="1ms"/>
|
| 34 |
+
<Line id="n" quantity="hhpop[0]/bioPhys1/membraneProperties/kChans/kChan/n/q" scale="1" color="#0000ff" timeScale="1ms"/>
|
| 35 |
+
</Display>
|
| 36 |
+
|
| 37 |
+
<Display id="d3" title="Hodgkin-Huxley Neuron: Current" timeScale="1ms" xmin="-20" xmax="470" ymin="-10" ymax="10">
|
| 38 |
+
<Line id="I_na" quantity="hhpop[0]/bioPhys1/membraneProperties/naChans/iDensity" scale="1" color="#00ffff" timeScale="1ms"/>
|
| 39 |
+
<Line id="I_k" quantity="hhpop[0]/bioPhys1/membraneProperties/kChans/iDensity" scale="1" color="#ffff00" timeScale="1ms"/>
|
| 40 |
+
<Line id="I_l" quantity="hhpop[0]/bioPhys1/membraneProperties/leak/iDensity" scale="1" color="#ff00ff" timeScale="1ms"/>
|
| 41 |
+
</Display>
|
| 42 |
+
|
| 43 |
+
<Display id="d4" title="Hodgkin-Huxley Neuron: I_inj (nA)" timeScale="1ms" xmin="-20" xmax="470" ymin="-0.01" ymax="0.4">
|
| 44 |
+
<Line id="I_inj1" quantity="hhpop[0]/pulseGen1/i" scale="1nA" color="#ffffff" timeScale="1ms"/>
|
| 45 |
+
<Line id="I_inj2" quantity="hhpop[0]/pulseGen2/i" scale="1nA" color="#000000" timeScale="1ms"/>
|
| 46 |
+
</Display>
|
| 47 |
+
|
| 48 |
+
<!-- Saved the membrane potential to file: hh_v.dat -->
|
| 49 |
+
<OutputFile id="of0" fileName="hh_v.dat">
|
| 50 |
+
<OutputColumn id="v" quantity="hhpop[0]/v"/>
|
| 51 |
+
</OutputFile>
|
| 52 |
+
|
| 53 |
+
<!-- for plotting results in JupyterNotebook -->
|
| 54 |
+
<OutputFile id="of1" fileName="hh_forJupyterNotebook.dat">
|
| 55 |
+
<OutputColumn id="v" quantity="hhpop[0]/v"/>
|
| 56 |
+
<OutputColumn id="m" quantity="hhpop[0]/bioPhys1/membraneProperties/naChans/naChan/m/q"/>
|
| 57 |
+
<OutputColumn id="h" quantity="hhpop[0]/bioPhys1/membraneProperties/naChans/naChan/h/q"/>
|
| 58 |
+
<OutputColumn id="n" quantity="hhpop[0]/bioPhys1/membraneProperties/kChans/kChan/n/q"/>
|
| 59 |
+
<OutputColumn id="I_na" quantity="hhpop[0]/bioPhys1/membraneProperties/naChans/iDensity"/>
|
| 60 |
+
<OutputColumn id="I_k" quantity="hhpop[0]/bioPhys1/membraneProperties/kChans/iDensity"/>
|
| 61 |
+
<OutputColumn id="I_l" quantity="hhpop[0]/bioPhys1/membraneProperties/leak/iDensity"/>
|
| 62 |
+
<OutputColumn id="I_inj1" quantity="hhpop[0]/pulseGen1/i"/>
|
| 63 |
+
<OutputColumn id="I_inj2" quantity="hhpop[0]/pulseGen2/i"/>
|
| 64 |
+
</OutputFile>
|
| 65 |
+
|
| 66 |
+
</Simulation>
|
| 67 |
+
|
| 68 |
+
|
| 69 |
+
</Lems>
|
Tutorial/Source/LEMS_HH_SingleAP.xml
ADDED
|
@@ -0,0 +1,41 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<Lems>
|
| 2 |
+
|
| 3 |
+
<!-- Example with Simple Hodgkin-Huxley cell specifying segment details-->
|
| 4 |
+
|
| 5 |
+
<!-- This is a file which can be read and executed by the LEMS Interpreter.
|
| 6 |
+
It imports the LEMS definitions of the core NeuroML 2 Components,
|
| 7 |
+
imports in "pure" NeuroML 2 and contains some LEMS elements for running
|
| 8 |
+
a simulation -->
|
| 9 |
+
|
| 10 |
+
|
| 11 |
+
<Target component="sim1"/>
|
| 12 |
+
|
| 13 |
+
<Include file="Cells.xml"/>
|
| 14 |
+
<Include file="Networks.xml"/>
|
| 15 |
+
<Include file="Simulation.xml"/>
|
| 16 |
+
|
| 17 |
+
|
| 18 |
+
<Include file="HHCellSingleAP.net.nml"/>
|
| 19 |
+
|
| 20 |
+
<Include file="hhcell.cell.nml"/>
|
| 21 |
+
<Include file="passiveChan.channel.nml"/>
|
| 22 |
+
<Include file="naChan.channel.nml"/>
|
| 23 |
+
<Include file="kChan.channel.nml"/>
|
| 24 |
+
|
| 25 |
+
|
| 26 |
+
<Simulation id="sim1" length="50ms" step="0.001ms" target="HHCellNetwork">
|
| 27 |
+
|
| 28 |
+
<Display id="d1" title="Hodgkin-Huxley Neuron: V (mV)" timeScale="1ms" xmin="-5" xmax="55" ymin="-90" ymax="50">
|
| 29 |
+
<Line id="V" quantity="hhpop/0/hhcell/v" scale="1mV" color="#000000" timeScale="1ms"/>
|
| 30 |
+
</Display>
|
| 31 |
+
|
| 32 |
+
|
| 33 |
+
<OutputFile id="of0" fileName="hh_v.dat">
|
| 34 |
+
<OutputColumn id="v" quantity="hhpop/0/hhcell/v"/>
|
| 35 |
+
</OutputFile>
|
| 36 |
+
|
| 37 |
+
|
| 38 |
+
</Simulation>
|
| 39 |
+
|
| 40 |
+
|
| 41 |
+
</Lems>
|
Tutorial/Source/LEMS_HH_VClamp.xml
ADDED
|
@@ -0,0 +1,67 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<Lems>
|
| 2 |
+
|
| 3 |
+
<!-- Example with Simple Hodgkin-Huxley cell specifying segment details-->
|
| 4 |
+
|
| 5 |
+
<!-- This is a file which can be read and executed by the LEMS Interpreter.
|
| 6 |
+
It imports the LEMS definitions of the core NeuroML 2 Components,
|
| 7 |
+
imports in "pure" NeuroML 2 and contains some LEMS elements for running
|
| 8 |
+
a simulation -->
|
| 9 |
+
|
| 10 |
+
|
| 11 |
+
<Target component="sim1"/>
|
| 12 |
+
|
| 13 |
+
<Include file="Cells.xml"/>
|
| 14 |
+
<Include file="Networks.xml"/>
|
| 15 |
+
<Include file="Simulation.xml"/>
|
| 16 |
+
|
| 17 |
+
<Include file="vclamp.xml"/>
|
| 18 |
+
|
| 19 |
+
|
| 20 |
+
<Include file="HHCellVClamp.net.nml"/>
|
| 21 |
+
<Include file="hhcell.cell.nml"/>
|
| 22 |
+
<Include file="passiveChan.channel.nml"/>
|
| 23 |
+
<Include file="naChan.channel.nml"/>
|
| 24 |
+
<Include file="kChan.channel.nml"/>
|
| 25 |
+
|
| 26 |
+
|
| 27 |
+
<Simulation id="sim1" length="50ms" step="0.0001ms" target="HHCellNetwork">
|
| 28 |
+
|
| 29 |
+
<Display id="d1" title="Hodgkin-Huxley Neuron: V (mV)" timeScale="1ms" xmin="-5" xmax="55" ymin="-90" ymax="50">
|
| 30 |
+
<Line id="V" quantity="hhpop[0]/v" scale="1mV" color="#000000" timeScale="1ms"/>
|
| 31 |
+
</Display>
|
| 32 |
+
|
| 33 |
+
<Display id="d2" title="Hodgkin-Huxley Neuron: Gating Variables" timeScale="1ms" xmin="-5" xmax="55" ymin="-0.1" ymax="1.1">
|
| 34 |
+
<Line id="m" quantity="hhpop[0]/bioPhys1/membraneProperties/naChans/naChan/m/q" scale="1" color="#ff0000" timeScale="1ms"/>
|
| 35 |
+
<Line id="h" quantity="hhpop[0]/bioPhys1/membraneProperties/naChans/naChan/h/q" scale="1" color="#00dd00" timeScale="1ms"/>
|
| 36 |
+
<Line id="n" quantity="hhpop[0]/bioPhys1/membraneProperties/kChans/kChan/n/q" scale="1" color="#0000ff" timeScale="1ms"/>
|
| 37 |
+
</Display>
|
| 38 |
+
|
| 39 |
+
<Display id="d3" title="Hodgkin-Huxley Neuron: Current" timeScale="1ms" xmin="-5" xmax="55" ymin="-30" ymax="20">
|
| 40 |
+
<Line id="I_na" quantity="hhpop[0]/bioPhys1/membraneProperties/naChans/iDensity" scale="1" color="#00ffff" timeScale="1ms"/>
|
| 41 |
+
<Line id="I_k" quantity="hhpop[0]/bioPhys1/membraneProperties/kChans/iDensity" scale="1" color="#ffff00" timeScale="1ms"/>
|
| 42 |
+
<Line id="I_l" quantity="hhpop[0]/bioPhys1/membraneProperties/leak/iDensity" scale="1" color="#ff00ff" timeScale="1ms"/>
|
| 43 |
+
</Display>
|
| 44 |
+
|
| 45 |
+
<Display id="d4" title="Hodgkin-Huxley Neuron: I_vClamp (nA)" timeScale="1ms" xmin="-5" xmax="55" ymin="-15" ymax="25">
|
| 46 |
+
<Line id="I_vClamp" quantity="hhpop[0]/vClamp/i" scale="1nA" color="#ffffff" timeScale="1ms"/>
|
| 47 |
+
</Display>
|
| 48 |
+
|
| 49 |
+
<!-- Saved the membrane potential to file: hh_v.dat -->
|
| 50 |
+
<OutputFile id="of0" fileName="hh_v.dat">
|
| 51 |
+
<OutputColumn id="v" quantity="hhpop[0]/v"/>
|
| 52 |
+
</OutputFile>
|
| 53 |
+
|
| 54 |
+
<OutputFile id="of1" fileName="hh_forJupyterNotebook.dat">
|
| 55 |
+
<OutputColumn id="v" quantity="hhpop[0]/v" />
|
| 56 |
+
<OutputColumn id="m" quantity="hhpop[0]/bioPhys1/membraneProperties/naChans/naChan/m/q" />
|
| 57 |
+
<OutputColumn id="h" quantity="hhpop[0]/bioPhys1/membraneProperties/naChans/naChan/h/q" />
|
| 58 |
+
<OutputColumn id="n" quantity="hhpop[0]/bioPhys1/membraneProperties/kChans/kChan/n/q" />
|
| 59 |
+
<OutputColumn id="I_na" quantity="hhpop[0]/bioPhys1/membraneProperties/naChans/iDensity" />
|
| 60 |
+
<OutputColumn id="I_k" quantity="hhpop[0]/bioPhys1/membraneProperties/kChans/iDensity" />
|
| 61 |
+
<OutputColumn id="I_l" quantity="hhpop[0]/bioPhys1/membraneProperties/leak/iDensity" />
|
| 62 |
+
<OutputColumn id="I_vClamp" quantity="hhpop[0]/vClamp/i" />
|
| 63 |
+
</OutputFile>
|
| 64 |
+
</Simulation>
|
| 65 |
+
|
| 66 |
+
|
| 67 |
+
</Lems>
|
Tutorial/Source/__init__.py
ADDED
|
@@ -0,0 +1,3 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
__all__ = ['HodgkinHuxley']
|
| 2 |
+
|
| 3 |
+
from Source.HodgkinHuxley import HodgkinHuxley
|
Tutorial/Source/hhcell.cell.nml
ADDED
|
@@ -0,0 +1,51 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<?xml version="1.0" encoding="UTF-8"?>
|
| 2 |
+
|
| 3 |
+
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2"
|
| 4 |
+
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
|
| 5 |
+
xsi:schemaLocation="http://www.neuroml.org/schema/neuroml2 https://raw.githubusercontent.com/NeuroML/NeuroML2/master/Schemas/NeuroML2/NeuroML_v2.2.xsd"
|
| 6 |
+
id="hhcell">
|
| 7 |
+
|
| 8 |
+
<include href="passiveChan.channel.nml"/> <!-- Include the channel definitions -->
|
| 9 |
+
<include href="naChan.channel.nml"/>
|
| 10 |
+
<include href="kChan.channel.nml"/>
|
| 11 |
+
|
| 12 |
+
<cell id="hhcell">
|
| 13 |
+
|
| 14 |
+
<notes>Conductance based cell model NeuroML2 format: standard Hodgkin Huxley model cell with Na, K and passive conductances</notes>
|
| 15 |
+
|
| 16 |
+
<morphology id="morphology">
|
| 17 |
+
<segment id="0" name="soma">
|
| 18 |
+
<proximal x="0" y="0" z="0" diameter="17.841242"/> <!--Gives a convenient surface area of 1000.0 um^2-->
|
| 19 |
+
<distal x="0" y="0" z="0" diameter="17.841242"/>
|
| 20 |
+
</segment>
|
| 21 |
+
|
| 22 |
+
<segmentGroup id="soma_group">
|
| 23 |
+
<member segment="0"/>
|
| 24 |
+
</segmentGroup>
|
| 25 |
+
|
| 26 |
+
</morphology>
|
| 27 |
+
|
| 28 |
+
<biophysicalProperties id="bioPhys1">
|
| 29 |
+
|
| 30 |
+
<membraneProperties>
|
| 31 |
+
|
| 32 |
+
<channelDensity id="leak" ionChannel="passiveChan" condDensity="0.3 mS_per_cm2" erev="-54.387mV" ion="non_specific"/>
|
| 33 |
+
<channelDensity id="naChans" ionChannel="naChan" condDensity="120.0 mS_per_cm2" erev="50.0 mV" ion="na"/>
|
| 34 |
+
<channelDensity id="kChans" ionChannel="kChan" condDensity="36 mS_per_cm2" erev="-77mV" ion="k"/>
|
| 35 |
+
|
| 36 |
+
<spikeThresh value="-20mV"/>
|
| 37 |
+
<specificCapacitance value="1.0 uF_per_cm2"/>
|
| 38 |
+
<initMembPotential value="-65mV"/>
|
| 39 |
+
|
| 40 |
+
</membraneProperties>
|
| 41 |
+
|
| 42 |
+
<intracellularProperties>
|
| 43 |
+
<resistivity value="0.03 kohm_cm"/> <!-- Note: not used in single compartment simulations -->
|
| 44 |
+
</intracellularProperties>
|
| 45 |
+
|
| 46 |
+
</biophysicalProperties>
|
| 47 |
+
|
| 48 |
+
</cell>
|
| 49 |
+
|
| 50 |
+
|
| 51 |
+
</neuroml>
|
Tutorial/Source/kChan.channel.nml
ADDED
|
@@ -0,0 +1,22 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<?xml version="1.0" encoding="UTF-8"?>
|
| 2 |
+
|
| 3 |
+
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2"
|
| 4 |
+
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
|
| 5 |
+
xsi:schemaLocation="http://www.neuroml.org/schema/neuroml2 https://raw.githubusercontent.com/NeuroML/NeuroML2/master/Schemas/NeuroML2/NeuroML_v2beta3.xsd"
|
| 6 |
+
id="kChan">
|
| 7 |
+
|
| 8 |
+
|
| 9 |
+
<ionChannelHH id="kChan" conductance="10pS" species="k">
|
| 10 |
+
|
| 11 |
+
<notes>Single ion channel in NeuroML2 format: standard Potassium channel from the Hodgkin Huxley model</notes>
|
| 12 |
+
|
| 13 |
+
<gateHHrates id="n" instances="4">
|
| 14 |
+
<forwardRate type="HHExpLinearRate" rate="0.1per_ms" midpoint="-55mV" scale="10mV"/>
|
| 15 |
+
<reverseRate type="HHExpRate" rate="0.125per_ms" midpoint="-65mV" scale="-80mV"/>
|
| 16 |
+
</gateHHrates>
|
| 17 |
+
|
| 18 |
+
</ionChannelHH>
|
| 19 |
+
|
| 20 |
+
|
| 21 |
+
</neuroml>
|
| 22 |
+
|
Tutorial/Source/naChan.channel.nml
ADDED
|
@@ -0,0 +1,27 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<?xml version="1.0" encoding="UTF-8"?>
|
| 2 |
+
|
| 3 |
+
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2"
|
| 4 |
+
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
|
| 5 |
+
xsi:schemaLocation="http://www.neuroml.org/schema/neuroml2 https://raw.githubusercontent.com/NeuroML/NeuroML2/master/Schemas/NeuroML2/NeuroML_v2beta3.xsd"
|
| 6 |
+
id="naChan">
|
| 7 |
+
|
| 8 |
+
|
| 9 |
+
<ionChannelHH id="naChan" conductance="10pS" species="na">
|
| 10 |
+
|
| 11 |
+
<notes>Single ion channel in NeuroML2 format: standard Sodium channel from the Hodgkin Huxley model</notes>
|
| 12 |
+
|
| 13 |
+
<gateHHrates id="m" instances="3">
|
| 14 |
+
<forwardRate type="HHExpLinearRate" rate="1per_ms" midpoint="-40mV" scale="10mV"/>
|
| 15 |
+
<reverseRate type="HHExpRate" rate="4per_ms" midpoint="-65mV" scale="-18mV"/>
|
| 16 |
+
</gateHHrates>
|
| 17 |
+
|
| 18 |
+
<gateHHrates id="h" instances="1">
|
| 19 |
+
<forwardRate type="HHExpRate" rate="0.07per_ms" midpoint="-65mV" scale="-20mV"/>
|
| 20 |
+
<reverseRate type="HHSigmoidRate" rate="1per_ms" midpoint="-35mV" scale="10mV"/>
|
| 21 |
+
</gateHHrates>
|
| 22 |
+
|
| 23 |
+
</ionChannelHH>
|
| 24 |
+
|
| 25 |
+
|
| 26 |
+
</neuroml>
|
| 27 |
+
|
Tutorial/Source/nmllite/HH.py
ADDED
|
@@ -0,0 +1,82 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
from neuromllite import Network, Cell, InputSource, Population, Synapse
|
| 2 |
+
from neuromllite import Projection, RandomConnectivity, Input, Simulation
|
| 3 |
+
import sys
|
| 4 |
+
|
| 5 |
+
################################################################################
|
| 6 |
+
### Build new network
|
| 7 |
+
|
| 8 |
+
net = Network(id="HHTest")
|
| 9 |
+
net.notes = "Example HH cell"
|
| 10 |
+
net.parameters = {"N": 1}
|
| 11 |
+
|
| 12 |
+
|
| 13 |
+
cell = Cell(id="hhcell", neuroml2_source_file="../hhcell.cell.nml")
|
| 14 |
+
cell.parameters = {}
|
| 15 |
+
|
| 16 |
+
params = {
|
| 17 |
+
}
|
| 18 |
+
|
| 19 |
+
|
| 20 |
+
for p in params:
|
| 21 |
+
cell.parameters[p] = p
|
| 22 |
+
net.parameters[p] = params[p]
|
| 23 |
+
|
| 24 |
+
net.cells.append(cell)
|
| 25 |
+
|
| 26 |
+
|
| 27 |
+
pop = Population(
|
| 28 |
+
id="hhPop", size="1", component=cell.id, properties={"color": ".7 0 0"}
|
| 29 |
+
)
|
| 30 |
+
net.populations.append(pop)
|
| 31 |
+
|
| 32 |
+
net.parameters["delay"] = "5ms"
|
| 33 |
+
net.parameters["stim_amp"] = "0.05nA"
|
| 34 |
+
net.parameters["duration"] = "25ms"
|
| 35 |
+
input_source = InputSource(
|
| 36 |
+
id="iclamp_0",
|
| 37 |
+
neuroml2_input="pulseGenerator",
|
| 38 |
+
parameters={"amplitude": "stim_amp", "delay": "delay", "duration": "duration"},
|
| 39 |
+
)
|
| 40 |
+
net.input_sources.append(input_source)
|
| 41 |
+
|
| 42 |
+
net.inputs.append(
|
| 43 |
+
Input(
|
| 44 |
+
id="stim",
|
| 45 |
+
input_source=input_source.id,
|
| 46 |
+
population=pop.id,
|
| 47 |
+
percentage=100,
|
| 48 |
+
weight=1,
|
| 49 |
+
)
|
| 50 |
+
)
|
| 51 |
+
|
| 52 |
+
print(net)
|
| 53 |
+
print(net.to_json())
|
| 54 |
+
net_yaml_file = net.to_yaml_file("%s.nmllite.yaml" % net.id)
|
| 55 |
+
net_json_file = net.to_json_file("%s.nmllite.json" % net.id)
|
| 56 |
+
|
| 57 |
+
|
| 58 |
+
################################################################################
|
| 59 |
+
### Build Simulation object & save as JSON
|
| 60 |
+
|
| 61 |
+
record_variables = {"v": {"all": "*"}}
|
| 62 |
+
|
| 63 |
+
sim = Simulation(
|
| 64 |
+
id="Sim%s" % net.id,
|
| 65 |
+
network=net_yaml_file,
|
| 66 |
+
duration="50",
|
| 67 |
+
dt="0.025",
|
| 68 |
+
record_variables=record_variables,
|
| 69 |
+
)
|
| 70 |
+
|
| 71 |
+
sim.to_yaml_file("%s.yaml" % sim.id)
|
| 72 |
+
sim.network = net_json_file
|
| 73 |
+
sim.to_json_file("%s.json" % sim.id)
|
| 74 |
+
|
| 75 |
+
|
| 76 |
+
################################################################################
|
| 77 |
+
### Run in some simulators
|
| 78 |
+
|
| 79 |
+
from neuromllite.NetworkGenerator import check_to_generate_or_run
|
| 80 |
+
import sys
|
| 81 |
+
|
| 82 |
+
check_to_generate_or_run(sys.argv, sim)
|
Tutorial/Source/nmllite/HHTest.gv
ADDED
|
@@ -0,0 +1,7 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
digraph HHTest {
|
| 2 |
+
node [color="#b20000" fontcolor="#ffffff" shape=ellipse style=filled]
|
| 3 |
+
hhPop [label=<hhPop<br/><i>1 cell</i>>]
|
| 4 |
+
node [color="#444444" fontcolor="#444444" style=""]
|
| 5 |
+
stim [label=<stim<br/><i>1 input</i>>]
|
| 6 |
+
stim -> hhPop [arrowhead=empty]
|
| 7 |
+
}
|
Tutorial/Source/nmllite/HHTest.gv.png
ADDED
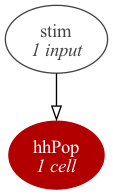
|
Tutorial/Source/nmllite/HHTest.mdf.json
ADDED
|
@@ -0,0 +1,70 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"HHTest": {
|
| 3 |
+
"format": "ModECI MDF v0.3",
|
| 4 |
+
"graphs": {
|
| 5 |
+
"HHTest": {
|
| 6 |
+
"notes": "Example HH cell",
|
| 7 |
+
"nodes": {
|
| 8 |
+
"hhPop_0": {
|
| 9 |
+
"parameters": {},
|
| 10 |
+
"input_ports": {},
|
| 11 |
+
"output_ports": {},
|
| 12 |
+
"notes": "Cell: [Cell(notes=None, id='hhcell', parameters={}, neuroml2_source_file='../hhcell.cell.nml', lems_source_file=None, neuroml2_cell=None, pynn_cell=None, arbor_cell=None, bindsnet_node=None)] is defined in None and in Lems is: Component, id: hhcell, type: None,\n parameters: {}\n parent: None\n"
|
| 13 |
+
},
|
| 14 |
+
"Input_stim_0": {
|
| 15 |
+
"parameters": {
|
| 16 |
+
"amplitude": {
|
| 17 |
+
"value": 5.000000000000001e-11
|
| 18 |
+
},
|
| 19 |
+
"delay": {
|
| 20 |
+
"value": 0.005
|
| 21 |
+
},
|
| 22 |
+
"duration": {
|
| 23 |
+
"value": 0.025
|
| 24 |
+
},
|
| 25 |
+
"i": {
|
| 26 |
+
"conditions": {
|
| 27 |
+
"condition_0": {
|
| 28 |
+
"test": "t < delay",
|
| 29 |
+
"value": "0"
|
| 30 |
+
},
|
| 31 |
+
"condition_1": {
|
| 32 |
+
"test": "t >= delay and t < duration + delay",
|
| 33 |
+
"value": "weight * amplitude"
|
| 34 |
+
},
|
| 35 |
+
"condition_2": {
|
| 36 |
+
"test": "t >= duration + delay",
|
| 37 |
+
"value": "0"
|
| 38 |
+
}
|
| 39 |
+
}
|
| 40 |
+
},
|
| 41 |
+
"t": {
|
| 42 |
+
"default_initial_value": 0,
|
| 43 |
+
"time_derivative": "1"
|
| 44 |
+
},
|
| 45 |
+
"weight": {
|
| 46 |
+
"value": 1
|
| 47 |
+
}
|
| 48 |
+
},
|
| 49 |
+
"input_ports": {},
|
| 50 |
+
"output_ports": {
|
| 51 |
+
"i": {
|
| 52 |
+
"value": "i"
|
| 53 |
+
}
|
| 54 |
+
},
|
| 55 |
+
"notes": "Cell: [InputSource(notes=None, id='iclamp_0', parameters={'amplitude': 'stim_amp', 'delay': 'delay', 'duration': 'duration'}, neuroml2_source_file=None, neuroml2_input='pulseGenerator', lems_source_file=None, pynn_input=None)] is defined in None and in Lems is: Component, id: iclamp_0, type: pulseGenerator,\n parameters: {'amplitude': '0.05nA', 'delay': '5ms', 'duration': '25ms'}\n parent: None\n"
|
| 56 |
+
}
|
| 57 |
+
},
|
| 58 |
+
"edges": {
|
| 59 |
+
"Edge Input_stim_0 to hhPop_0": {
|
| 60 |
+
"name": "Edge Input_stim_0 to hhPop_0",
|
| 61 |
+
"sender_port": "i",
|
| 62 |
+
"receiver_port": "synapses_i",
|
| 63 |
+
"sender": "Input_stim_0",
|
| 64 |
+
"receiver": "hhPop_0"
|
| 65 |
+
}
|
| 66 |
+
}
|
| 67 |
+
}
|
| 68 |
+
}
|
| 69 |
+
}
|
| 70 |
+
}
|
Tutorial/Source/nmllite/HHTest.mdf.yaml
ADDED
|
@@ -0,0 +1,57 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
HHTest:
|
| 2 |
+
format: ModECI MDF v0.3
|
| 3 |
+
graphs:
|
| 4 |
+
HHTest:
|
| 5 |
+
notes: Example HH cell
|
| 6 |
+
nodes:
|
| 7 |
+
hhPop_0:
|
| 8 |
+
parameters: {}
|
| 9 |
+
input_ports: {}
|
| 10 |
+
output_ports: {}
|
| 11 |
+
notes: "Cell: [Cell(notes=None, id='hhcell', parameters={}, neuroml2_source_file='../hhcell.cell.nml',\
|
| 12 |
+
\ lems_source_file=None, neuroml2_cell=None, pynn_cell=None,\
|
| 13 |
+
\ arbor_cell=None, bindsnet_node=None)] is defined in None\
|
| 14 |
+
\ and in Lems is: Component, id: hhcell, type: None,\n parameters:\
|
| 15 |
+
\ {}\n parent: None\n"
|
| 16 |
+
Input_stim_0:
|
| 17 |
+
parameters:
|
| 18 |
+
amplitude:
|
| 19 |
+
value: 5.000000000000001e-11
|
| 20 |
+
delay:
|
| 21 |
+
value: 0.005
|
| 22 |
+
duration:
|
| 23 |
+
value: 0.025
|
| 24 |
+
i:
|
| 25 |
+
conditions:
|
| 26 |
+
condition_0:
|
| 27 |
+
test: t < delay
|
| 28 |
+
value: '0'
|
| 29 |
+
condition_1:
|
| 30 |
+
test: t >= delay and t < duration + delay
|
| 31 |
+
value: weight * amplitude
|
| 32 |
+
condition_2:
|
| 33 |
+
test: t >= duration + delay
|
| 34 |
+
value: '0'
|
| 35 |
+
t:
|
| 36 |
+
default_initial_value: 0
|
| 37 |
+
time_derivative: '1'
|
| 38 |
+
weight:
|
| 39 |
+
value: 1
|
| 40 |
+
input_ports: {}
|
| 41 |
+
output_ports:
|
| 42 |
+
i:
|
| 43 |
+
value: i
|
| 44 |
+
notes: "Cell: [InputSource(notes=None, id='iclamp_0', parameters={'amplitude':\
|
| 45 |
+
\ 'stim_amp', 'delay': 'delay', 'duration': 'duration'}, neuroml2_source_file=None,\
|
| 46 |
+
\ neuroml2_input='pulseGenerator', lems_source_file=None,\
|
| 47 |
+
\ pynn_input=None)] is defined in None and in Lems is: Component,\
|
| 48 |
+
\ id: iclamp_0, type: pulseGenerator,\n parameters: {'amplitude':\
|
| 49 |
+
\ '0.05nA', 'delay': '5ms', 'duration': '25ms'}\n parent:\
|
| 50 |
+
\ None\n"
|
| 51 |
+
edges:
|
| 52 |
+
Edge Input_stim_0 to hhPop_0:
|
| 53 |
+
name: Edge Input_stim_0 to hhPop_0
|
| 54 |
+
sender_port: i
|
| 55 |
+
receiver_port: synapses_i
|
| 56 |
+
sender: Input_stim_0
|
| 57 |
+
receiver: hhPop_0
|
Tutorial/Source/nmllite/HHTest.net.nml
ADDED
|
@@ -0,0 +1,21 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2" xmlns:xs="http://www.w3.org/2001/XMLSchema" xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance" xsi:schemaLocation="http://www.neuroml.org/schema/neuroml2 https://raw.github.com/NeuroML/NeuroML2/development/Schemas/NeuroML2/NeuroML_v2.3.xsd" id="HHTest">
|
| 2 |
+
<notes>Generated by NeuroMLlite v0.5.8
|
| 3 |
+
Generated network: HHTest
|
| 4 |
+
Generation seed: 1234
|
| 5 |
+
NeuroMLlite parameters:
|
| 6 |
+
N = 10.0
|
| 7 |
+
delay = 5ms
|
| 8 |
+
duration = 25ms
|
| 9 |
+
stim_amp = -0.5nA</notes>
|
| 10 |
+
<include href="/Users/padraig/neuroConstruct/osb/generic/hodgkin_huxley_tutorial/Tutorial/Source/hhcell.cell.nml"/>
|
| 11 |
+
<pulseGenerator id="iclamp_0" delay="5ms" duration="25ms" amplitude="-0.5nA"/>
|
| 12 |
+
<network id="HHTest">
|
| 13 |
+
<notes>Example HH cell</notes>
|
| 14 |
+
<property tag="recommended_dt_ms" value="0.025"/>
|
| 15 |
+
<property tag="recommended_duration_ms" value="50.0"/>
|
| 16 |
+
<population id="hhPop" component="hhcell" size="1"/>
|
| 17 |
+
<inputList id="stim" population="hhPop" component="iclamp_0">
|
| 18 |
+
<input id="0" target="../hhPop[0]" destination="synapses"/>
|
| 19 |
+
</inputList>
|
| 20 |
+
</network>
|
| 21 |
+
</neuroml>
|
Tutorial/Source/nmllite/HHTest.net.png
ADDED
|
Tutorial/Source/nmllite/HHTest.nmllite.json
ADDED
|
@@ -0,0 +1,45 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"HHTest": {
|
| 3 |
+
"version": "NeuroMLlite v0.5.8",
|
| 4 |
+
"notes": "Example HH cell",
|
| 5 |
+
"parameters": {
|
| 6 |
+
"N": 1,
|
| 7 |
+
"delay": "5ms",
|
| 8 |
+
"stim_amp": "0.05nA",
|
| 9 |
+
"duration": "25ms"
|
| 10 |
+
},
|
| 11 |
+
"cells": {
|
| 12 |
+
"hhcell": {
|
| 13 |
+
"parameters": {},
|
| 14 |
+
"neuroml2_source_file": "../hhcell.cell.nml"
|
| 15 |
+
}
|
| 16 |
+
},
|
| 17 |
+
"input_sources": {
|
| 18 |
+
"iclamp_0": {
|
| 19 |
+
"parameters": {
|
| 20 |
+
"amplitude": "stim_amp",
|
| 21 |
+
"delay": "delay",
|
| 22 |
+
"duration": "duration"
|
| 23 |
+
},
|
| 24 |
+
"neuroml2_input": "pulseGenerator"
|
| 25 |
+
}
|
| 26 |
+
},
|
| 27 |
+
"populations": {
|
| 28 |
+
"hhPop": {
|
| 29 |
+
"size": "1",
|
| 30 |
+
"component": "hhcell",
|
| 31 |
+
"properties": {
|
| 32 |
+
"color": ".7 0 0"
|
| 33 |
+
}
|
| 34 |
+
}
|
| 35 |
+
},
|
| 36 |
+
"inputs": {
|
| 37 |
+
"stim": {
|
| 38 |
+
"input_source": "iclamp_0",
|
| 39 |
+
"population": "hhPop",
|
| 40 |
+
"percentage": 100,
|
| 41 |
+
"weight": 1
|
| 42 |
+
}
|
| 43 |
+
}
|
| 44 |
+
}
|
| 45 |
+
}
|
Tutorial/Source/nmllite/HHTest.nmllite.yaml
ADDED
|
@@ -0,0 +1,31 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
HHTest:
|
| 2 |
+
version: NeuroMLlite v0.5.8
|
| 3 |
+
notes: Example HH cell
|
| 4 |
+
parameters:
|
| 5 |
+
N: 1
|
| 6 |
+
delay: 5ms
|
| 7 |
+
stim_amp: 0.05nA
|
| 8 |
+
duration: 25ms
|
| 9 |
+
cells:
|
| 10 |
+
hhcell:
|
| 11 |
+
parameters: {}
|
| 12 |
+
neuroml2_source_file: ../hhcell.cell.nml
|
| 13 |
+
input_sources:
|
| 14 |
+
iclamp_0:
|
| 15 |
+
parameters:
|
| 16 |
+
amplitude: stim_amp
|
| 17 |
+
delay: delay
|
| 18 |
+
duration: duration
|
| 19 |
+
neuroml2_input: pulseGenerator
|
| 20 |
+
populations:
|
| 21 |
+
hhPop:
|
| 22 |
+
size: '1'
|
| 23 |
+
component: hhcell
|
| 24 |
+
properties:
|
| 25 |
+
color: .7 0 0
|
| 26 |
+
inputs:
|
| 27 |
+
stim:
|
| 28 |
+
input_source: iclamp_0
|
| 29 |
+
population: hhPop
|
| 30 |
+
percentage: 100
|
| 31 |
+
weight: 1
|
Tutorial/Source/nmllite/LEMS_SimHHTest.gv
ADDED
|
@@ -0,0 +1,254 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# GraphViz compliant export for:HHTest (Type: network)
|
| 2 |
+
|
| 3 |
+
digraph SimHHTest {
|
| 4 |
+
fontsize=10;
|
| 5 |
+
overlap=false;
|
| 6 |
+
|
| 7 |
+
|
| 8 |
+
subgraph cluster_network {
|
| 9 |
+
style=filled;
|
| 10 |
+
color="#D6eeEA";
|
| 11 |
+
node [style=filled,color=white];
|
| 12 |
+
label = "Network to be simulated";
|
| 13 |
+
|
| 14 |
+
node [shape=rectangle]; HHTest;
|
| 15 |
+
# Population hhPop contains components of: Component(id=hhcell type=cell)
|
| 16 |
+
node [shape=ellipse,color="white",fontcolor="black"]; hhPop;
|
| 17 |
+
HHTest -> hhPop [len=1.00, arrowhead=diamond]
|
| 18 |
+
|
| 19 |
+
}
|
| 20 |
+
|
| 21 |
+
subgraph cluster_comps {
|
| 22 |
+
style=filled;
|
| 23 |
+
color="#CCFFCC";
|
| 24 |
+
node [style=filled,color=white];
|
| 25 |
+
label = "Components";
|
| 26 |
+
|
| 27 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>hhcell (cell)</td></tr><tr><td><font color="#666666"></font></td></tr></table>>]; "hhcell (cell)";
|
| 28 |
+
|
| 29 |
+
"hhPop" -> "hhcell (cell)" [label="1",len=1.00, arrowhead=diamond]
|
| 30 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (0)</td></tr><tr><td><font color="#666666"></font></td></tr></table>>]; "null (0)";
|
| 31 |
+
|
| 32 |
+
"hhcell (cell)" -> "null (0)" [len=1.00, arrowhead=diamond]
|
| 33 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>morphology (morphology)</td></tr><tr><td><font color="#666666"></font></td></tr></table>>]; "morphology (morphology)";
|
| 34 |
+
|
| 35 |
+
"hhcell (cell)" -> "morphology (morphology)" [len=1.00, arrowhead=diamond]
|
| 36 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>soma (id = 0)</td></tr><tr><td><font color="#666666"></font></td></tr></table>>]; "soma (id = 0)";
|
| 37 |
+
|
| 38 |
+
"morphology (morphology)" -> "soma (id = 0)" [len=1.00, arrowhead=diamond]
|
| 39 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (1)</td></tr><tr><td><font color="#666666">x = 0, y = 0, z = 0, <br/>diameter = 17.841242, radius = 8.920621E-6 m, xLength = 0 m, <br/>yLength = 0 m, zLength = 0 m</font></td></tr></table>>]; "null (1)";
|
| 40 |
+
|
| 41 |
+
"soma (id = 0)" -> "null (1)" [len=1.00, arrowhead=diamond]
|
| 42 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (2)</td></tr><tr><td><font color="#666666">x = 0, y = 0, z = 0, <br/>diameter = 17.841242, radius = 8.920621E-6 m, xLength = 0 m, <br/>yLength = 0 m, zLength = 0 m</font></td></tr></table>>]; "null (2)";
|
| 43 |
+
|
| 44 |
+
"soma (id = 0)" -> "null (2)" [len=1.00, arrowhead=diamond]
|
| 45 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>soma_group (segmentGroup)</td></tr><tr><td><font color="#666666"></font></td></tr></table>>]; "soma_group (segmentGroup)";
|
| 46 |
+
|
| 47 |
+
"morphology (morphology)" -> "soma_group (segmentGroup)" [len=1.00, arrowhead=diamond]
|
| 48 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (3)</td></tr><tr><td><font color="#666666"></font></td></tr></table>>]; "null (3)";
|
| 49 |
+
|
| 50 |
+
"soma_group (segmentGroup)" -> "null (3)" [len=1.00, arrowhead=diamond]
|
| 51 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>bioPhys1 (biophysicalProperties)</td></tr><tr><td><font color="#666666"></font></td></tr></table>>]; "bioPhys1 (biophysicalProperties)";
|
| 52 |
+
|
| 53 |
+
"hhcell (cell)" -> "bioPhys1 (biophysicalProperties)" [len=1.00, arrowhead=diamond]
|
| 54 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (4)</td></tr><tr><td><font color="#666666"></font></td></tr></table>>]; "null (4)";
|
| 55 |
+
|
| 56 |
+
"bioPhys1 (biophysicalProperties)" -> "null (4)" [len=1.00, arrowhead=diamond]
|
| 57 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (5)</td></tr><tr><td><font color="#666666">value = -0.02 V</font></td></tr></table>>]; "null (5)";
|
| 58 |
+
|
| 59 |
+
"null (4)" -> "null (5)" [len=1.00, arrowhead=diamond]
|
| 60 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (6)</td></tr><tr><td><font color="#666666">value = -0.065 V</font></td></tr></table>>]; "null (6)";
|
| 61 |
+
|
| 62 |
+
"null (4)" -> "null (6)" [len=1.00, arrowhead=diamond]
|
| 63 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>leak (channelDensity)</td></tr><tr><td><font color="#666666">erev = -0.054387000000000005 V, condDensity = 3 kg^-1 m^-4 s^3 A^2</font></td></tr></table>>]; "leak (channelDensity)";
|
| 64 |
+
|
| 65 |
+
"null (4)" -> "leak (channelDensity)" [len=1.00, arrowhead=diamond]
|
| 66 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>passiveChan (ionChannelPassive)</td></tr><tr><td><font color="#666666">conductance = 1.0E-11 S</font></td></tr></table>>]; "passiveChan (ionChannelPassive)";
|
| 67 |
+
|
| 68 |
+
"leak (channelDensity)" -> "passiveChan (ionChannelPassive)" [len=1.00, arrowhead=diamond]
|
| 69 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (7)</td></tr><tr><td><font color="#666666"></font></td></tr></table>>]; "null (7)";
|
| 70 |
+
|
| 71 |
+
"passiveChan (ionChannelPassive)" -> "null (7)" [len=1.00, arrowhead=diamond]
|
| 72 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>naChans (channelDensity)</td></tr><tr><td><font color="#666666">erev = 0.05 V, condDensity = 1200 kg^-1 m^-4 s^3 A^2</font></td></tr></table>>]; "naChans (channelDensity)";
|
| 73 |
+
|
| 74 |
+
"null (4)" -> "naChans (channelDensity)" [len=1.00, arrowhead=diamond]
|
| 75 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>naChan (ionChannelHH)</td></tr><tr><td><font color="#666666">conductance = 1.0E-11 S</font></td></tr></table>>]; "naChan (ionChannelHH)";
|
| 76 |
+
|
| 77 |
+
"naChans (channelDensity)" -> "naChan (ionChannelHH)" [len=1.00, arrowhead=diamond]
|
| 78 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (8)</td></tr><tr><td><font color="#666666"></font></td></tr></table>>]; "null (8)";
|
| 79 |
+
|
| 80 |
+
"naChan (ionChannelHH)" -> "null (8)" [len=1.00, arrowhead=diamond]
|
| 81 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>m (gateHHrates)</td></tr><tr><td><font color="#666666">instances = 3</font></td></tr></table>>]; "m (gateHHrates)";
|
| 82 |
+
|
| 83 |
+
"naChan (ionChannelHH)" -> "m (gateHHrates)" [len=1.00, arrowhead=diamond]
|
| 84 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (9)</td></tr><tr><td><font color="#666666">rate = 1000 s^-1, midpoint = -0.04 V, scale = 0.01 V</font></td></tr></table>>]; "null (9)";
|
| 85 |
+
|
| 86 |
+
"m (gateHHrates)" -> "null (9)" [len=1.00, arrowhead=diamond]
|
| 87 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (10)</td></tr><tr><td><font color="#666666">rate = 4000 s^-1, midpoint = -0.065 V, scale = -0.018000000000000002 V</font></td></tr></table>>]; "null (10)";
|
| 88 |
+
|
| 89 |
+
"m (gateHHrates)" -> "null (10)" [len=1.00, arrowhead=diamond]
|
| 90 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>h (gateHHrates)</td></tr><tr><td><font color="#666666">instances = 1</font></td></tr></table>>]; "h (gateHHrates)";
|
| 91 |
+
|
| 92 |
+
"naChan (ionChannelHH)" -> "h (gateHHrates)" [len=1.00, arrowhead=diamond]
|
| 93 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (11)</td></tr><tr><td><font color="#666666">rate = 70 s^-1, midpoint = -0.065 V, scale = -0.02 V</font></td></tr></table>>]; "null (11)";
|
| 94 |
+
|
| 95 |
+
"h (gateHHrates)" -> "null (11)" [len=1.00, arrowhead=diamond]
|
| 96 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (12)</td></tr><tr><td><font color="#666666">rate = 1000 s^-1, midpoint = -0.035 V, scale = 0.01 V</font></td></tr></table>>]; "null (12)";
|
| 97 |
+
|
| 98 |
+
"h (gateHHrates)" -> "null (12)" [len=1.00, arrowhead=diamond]
|
| 99 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>kChans (channelDensity)</td></tr><tr><td><font color="#666666">erev = -0.077 V, condDensity = 360 kg^-1 m^-4 s^3 A^2</font></td></tr></table>>]; "kChans (channelDensity)";
|
| 100 |
+
|
| 101 |
+
"null (4)" -> "kChans (channelDensity)" [len=1.00, arrowhead=diamond]
|
| 102 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>kChan (ionChannelHH)</td></tr><tr><td><font color="#666666">conductance = 1.0E-11 S</font></td></tr></table>>]; "kChan (ionChannelHH)";
|
| 103 |
+
|
| 104 |
+
"kChans (channelDensity)" -> "kChan (ionChannelHH)" [len=1.00, arrowhead=diamond]
|
| 105 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (13)</td></tr><tr><td><font color="#666666"></font></td></tr></table>>]; "null (13)";
|
| 106 |
+
|
| 107 |
+
"kChan (ionChannelHH)" -> "null (13)" [len=1.00, arrowhead=diamond]
|
| 108 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>n (gateHHrates)</td></tr><tr><td><font color="#666666">instances = 4</font></td></tr></table>>]; "n (gateHHrates)";
|
| 109 |
+
|
| 110 |
+
"kChan (ionChannelHH)" -> "n (gateHHrates)" [len=1.00, arrowhead=diamond]
|
| 111 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (14)</td></tr><tr><td><font color="#666666">rate = 100 s^-1, midpoint = -0.055 V, scale = 0.01 V</font></td></tr></table>>]; "null (14)";
|
| 112 |
+
|
| 113 |
+
"n (gateHHrates)" -> "null (14)" [len=1.00, arrowhead=diamond]
|
| 114 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (15)</td></tr><tr><td><font color="#666666">rate = 125 s^-1, midpoint = -0.065 V, scale = -0.08 V</font></td></tr></table>>]; "null (15)";
|
| 115 |
+
|
| 116 |
+
"n (gateHHrates)" -> "null (15)" [len=1.00, arrowhead=diamond]
|
| 117 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (16)</td></tr><tr><td><font color="#666666">value = 0.01 kg^-1 m^-4 s^4 A^2</font></td></tr></table>>]; "null (16)";
|
| 118 |
+
|
| 119 |
+
"null (4)" -> "null (16)" [len=1.00, arrowhead=diamond]
|
| 120 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (17)</td></tr><tr><td><font color="#666666"></font></td></tr></table>>]; "null (17)";
|
| 121 |
+
|
| 122 |
+
"bioPhys1 (biophysicalProperties)" -> "null (17)" [len=1.00, arrowhead=diamond]
|
| 123 |
+
node [shape=ellipse label=<<table border="0" cellborder="0"><tr><td>null (18)</td></tr><tr><td><font color="#666666">value = 0.3 kg^2 m^2 s^-3 A^-2</font></td></tr></table>>]; "null (18)";
|
| 124 |
+
|
| 125 |
+
"null (17)" -> "null (18)" [len=1.00, arrowhead=diamond]
|
| 126 |
+
}
|
| 127 |
+
|
| 128 |
+
subgraph cluster_compTypes {
|
| 129 |
+
style=filled;
|
| 130 |
+
color="#D6E0EA";
|
| 131 |
+
node [style="rounded, filled",color=white];
|
| 132 |
+
label = "Component Types";
|
| 133 |
+
node [style="rounded, filled",color=white];
|
| 134 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>cell</td></tr><tr><td><font color="#B2C0D9">neuroLexId</font></td></tr><tr><td><font color="#FF9966">State vars: v (voltage), spiking </font></td></tr><tr><td><font color="#99CC00">initMembPot = biophysicalProperties/membraneProperties/initMembPotential/value</font></td></tr><tr><td><font color="#99CC00">thresh = biophysicalProperties/membraneProperties/spikeThresh/value</font></td></tr><tr><td><font color="#99CC00">surfaceArea = SUM OF: morphology/segments[*]/surfaceArea</font></td></tr><tr><td><font color="#99CC00">totSpecCap = biophysicalProperties/totSpecCap</font></td></tr><tr><td><font color="#99CC00">totCap = totSpecCap * surfaceArea </font></td></tr><tr><td><font color="#99CC00">iChannels = biophysicalProperties/membraneProperties/totChanCurrent</font></td></tr><tr><td><font color="#99CC00">iSyn = SUM OF: synapses[*]/i</font></td></tr><tr><td><font color="#99CC00">iCa = biophysicalProperties/membraneProperties/iCa</font></td></tr><tr><td><font color="#99CC00">caConc = biophysicalProperties/intracellularProperties/caConc</font></td></tr><tr><td><font color="#99CC00">caConcExt = biophysicalProperties/intracellularProperties/caConcExt</font></td></tr><tr><td><font color="#666633">v' = (iChannels + iSyn) / totCap</font></td></tr><tr><td><font color="#996633">IF v .gt. thresh AND spiking .lt. 0.5 THEN </font></td></tr><tr><td><font color="#996633">(spiking = 1) <br/>AND (EVENT: spike)</font></td></tr><tr><td><font color="#996633">IF v .lt. thresh THEN </font></td></tr><tr><td><font color="#996633">(spiking = 0)</font></td></tr><tr><td><font color="#666666">Exposures: spiking, iChannels (current), iSyn (current), <br/>totSpecCap (specificCapacitance), surfaceArea (area), iCa (current), <br/>caConc (concentration), caConcExt (concentration), v (voltage) </font></td></tr></table>>]; cell;
|
| 135 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseCellMembPot</td></tr><tr><td><font color="#666666">Exposures: v (voltage) </font></td></tr></table>>]; baseCellMembPot;
|
| 136 |
+
cell -> baseCellMembPot [len=1.00, arrowhead=onormal]
|
| 137 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseSpikingCell</td></tr></table>>]; baseSpikingCell;
|
| 138 |
+
baseCellMembPot -> baseSpikingCell [len=1.00, arrowhead=onormal]
|
| 139 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseCell</td></tr></table>>]; baseCell;
|
| 140 |
+
baseSpikingCell -> baseCell [len=1.00, arrowhead=onormal]
|
| 141 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseStandalone</td></tr></table>>]; baseStandalone;
|
| 142 |
+
baseCell -> baseStandalone [len=1.00, arrowhead=onormal]
|
| 143 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>notes</td></tr></table>>]; notes;
|
| 144 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>morphology</td></tr></table>>]; morphology;
|
| 145 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>segment</td></tr><tr><td><font color="#662211">Consts: LEN = 1 m </font></td></tr><tr><td><font color="#B2C0D9">name</font></td></tr><tr><td><font color="#99CC00">radDist = distal/radius</font></td></tr><tr><td><font color="#99CC00">dx = distal/xLength</font></td></tr><tr><td><font color="#99CC00">dy = distal/yLength</font></td></tr><tr><td><font color="#99CC00">dz = distal/zLength</font></td></tr><tr><td><font color="#99CC00">px = proximal/xLength</font></td></tr><tr><td><font color="#99CC00">py = proximal/yLength</font></td></tr><tr><td><font color="#99CC00">pz = proximal/zLength</font></td></tr><tr><td><font color="#99CC00">length = sqrt(((dx - px) * (dx - px) + (dy - py) * (dy - py) + (dz - pz) * (dz - pz))/(LEN * LEN)) * LEN</font></td></tr><tr><td><font color="#666666">Exposures: surfaceArea (area), radDist (length), length (length) </font></td></tr></table>>]; segment;
|
| 146 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>proximal</td></tr><tr><td><font color="#662211">Consts: MICRON = 1.0E-6 m </font></td></tr></table>>]; proximal;
|
| 147 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>point3DWithDiam</td></tr><tr><td><font color="#669999">Params: x, y, z, <br/>diameter </font></td></tr><tr><td><font color="#662211">Consts: MICRON = 1.0E-6 m </font></td></tr></table>>]; point3DWithDiam;
|
| 148 |
+
proximal -> point3DWithDiam [len=1.00, arrowhead=onormal]
|
| 149 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>distal</td></tr><tr><td><font color="#662211">Consts: MICRON = 1.0E-6 m </font></td></tr></table>>]; distal;
|
| 150 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>point3DWithDiam</td></tr><tr><td><font color="#669999">Params: x, y, z, <br/>diameter </font></td></tr><tr><td><font color="#662211">Consts: MICRON = 1.0E-6 m </font></td></tr></table>>]; point3DWithDiam;
|
| 151 |
+
distal -> point3DWithDiam [len=1.00, arrowhead=onormal]
|
| 152 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>segmentGroup</td></tr><tr><td><font color="#B2C0D9">neuroLexId</font></td></tr></table>>]; segmentGroup;
|
| 153 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>member</td></tr><tr><td><font color="#B2C0D9">segment</font></td></tr></table>>]; member;
|
| 154 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>biophysicalProperties</td></tr><tr><td><font color="#99CC00">totSpecCap = membraneProperties/totSpecCap</font></td></tr><tr><td><font color="#666666">Exposures: totSpecCap (specificCapacitance) </font></td></tr></table>>]; biophysicalProperties;
|
| 155 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>membraneProperties</td></tr><tr><td><font color="#666699">REQUIRES: surfaceArea (area)</font></td></tr><tr><td><font color="#99CC00">totSpecCap = SUM OF: specificCapacitances[*]/specCap</font></td></tr><tr><td><font color="#99CC00">totChanPopCurrent = SUM OF: populations[*]/i</font></td></tr><tr><td><font color="#99CC00">totChanDensCurrentDensity = SUM OF: channelDensities[*]/iDensity</font></td></tr><tr><td><font color="#99CC00">totChanCurrent = totChanPopCurrent + (totChanDensCurrentDensity * surfaceArea)</font></td></tr><tr><td><font color="#99CC00">totChanPopCurrentCa = SUM OF: populations[ion='ca']/i</font></td></tr><tr><td><font color="#99CC00">totChanDensCurrentDensityCa = SUM OF: channelDensities[ion='ca']/iDensity</font></td></tr><tr><td><font color="#99CC00">iCa = totChanPopCurrentCa + (totChanDensCurrentDensityCa * surfaceArea)</font></td></tr><tr><td><font color="#666666">Exposures: totChanCurrent (current), iCa (current), totSpecCap (specificCapacitance) </font></td></tr></table>>]; membraneProperties;
|
| 156 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>spikeThresh</td></tr><tr><td><font color="#669999">Params: value (voltage) </font></td></tr></table>>]; spikeThresh;
|
| 157 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>initMembPotential</td></tr><tr><td><font color="#669999">Params: value (voltage) </font></td></tr></table>>]; initMembPotential;
|
| 158 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>channelDensity</td></tr><tr><td><font color="#669999">Params: erev (voltage) </font></td></tr><tr><td><font color="#662211">Consts: vShift = 0 V </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#B2C0D9">segmentGroup</font></td></tr><tr><td><font color="#B2C0D9">ion</font></td></tr><tr><td><font color="#99CC00">channelf = ionChannel/fopen</font></td></tr><tr><td><font color="#99CC00">gDensity = condDensity * channelf</font></td></tr><tr><td><font color="#99CC00">iDensity = gDensity * (erev - v)</font></td></tr><tr><td><font color="#666666">Exposures: gDensity (conductanceDensity), iDensity (currentDensity) </font></td></tr></table>>]; channelDensity;
|
| 159 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseChannelDensityCond</td></tr><tr><td><font color="#669999">Params: condDensity (conductanceDensity) </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: gDensity (conductanceDensity), iDensity (currentDensity) </font></td></tr></table>>]; baseChannelDensityCond;
|
| 160 |
+
channelDensity -> baseChannelDensityCond [len=1.00, arrowhead=onormal]
|
| 161 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseChannelDensity</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: iDensity (currentDensity) </font></td></tr></table>>]; baseChannelDensity;
|
| 162 |
+
baseChannelDensityCond -> baseChannelDensity [len=1.00, arrowhead=onormal]
|
| 163 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>ionChannelPassive</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#B2C0D9">species</font></td></tr><tr><td><font color="#99CC00">fopen = 1</font></td></tr><tr><td><font color="#99CC00">g = conductance</font></td></tr><tr><td><font color="#666666">Exposures: g (conductance), fopen </font></td></tr></table>>]; ionChannelPassive;
|
| 164 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>ionChannel</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#99CC00">conductanceScale = PRODUCT OF: conductanceScaling[*]/factor</font></td></tr><tr><td><font color="#99CC00">fopen0 = PRODUCT OF: gates[*]/fcond</font></td></tr><tr><td><font color="#99CC00">fopen = conductanceScale * fopen0</font></td></tr><tr><td><font color="#99CC00">g = conductance * fopen</font></td></tr><tr><td><font color="#666666">Exposures: g (conductance), fopen </font></td></tr></table>>]; ionChannel;
|
| 165 |
+
ionChannelPassive -> ionChannel [len=1.00, arrowhead=onormal]
|
| 166 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>ionChannelHH</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#B2C0D9">species</font></td></tr><tr><td><font color="#99CC00">conductanceScale = PRODUCT OF: conductanceScaling[*]/factor</font></td></tr><tr><td><font color="#99CC00">fopen0 = PRODUCT OF: gates[*]/fcond</font></td></tr><tr><td><font color="#99CC00">fopen = conductanceScale * fopen0</font></td></tr><tr><td><font color="#99CC00">g = conductance * fopen</font></td></tr><tr><td><font color="#666666">Exposures: g (conductance), fopen </font></td></tr></table>>]; ionChannelHH;
|
| 167 |
+
ionChannel -> ionChannelHH [len=1.00, arrowhead=onormal]
|
| 168 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseIonChannel</td></tr><tr><td><font color="#669999">Params: conductance (conductance) </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#B2C0D9">neuroLexId</font></td></tr><tr><td><font color="#666666">Exposures: g (conductance), fopen </font></td></tr></table>>]; baseIonChannel;
|
| 169 |
+
ionChannelHH -> baseIonChannel [len=1.00, arrowhead=onormal]
|
| 170 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>notes</td></tr></table>>]; notes;
|
| 171 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>channelDensity</td></tr><tr><td><font color="#669999">Params: erev (voltage) </font></td></tr><tr><td><font color="#662211">Consts: vShift = 0 V </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#B2C0D9">segmentGroup</font></td></tr><tr><td><font color="#B2C0D9">ion</font></td></tr><tr><td><font color="#99CC00">channelf = ionChannel/fopen</font></td></tr><tr><td><font color="#99CC00">gDensity = condDensity * channelf</font></td></tr><tr><td><font color="#99CC00">iDensity = gDensity * (erev - v)</font></td></tr><tr><td><font color="#666666">Exposures: gDensity (conductanceDensity), iDensity (currentDensity) </font></td></tr></table>>]; channelDensity;
|
| 172 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseChannelDensityCond</td></tr><tr><td><font color="#669999">Params: condDensity (conductanceDensity) </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: gDensity (conductanceDensity), iDensity (currentDensity) </font></td></tr></table>>]; baseChannelDensityCond;
|
| 173 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseChannelDensity</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: iDensity (currentDensity) </font></td></tr></table>>]; baseChannelDensity;
|
| 174 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>ionChannelHH</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#B2C0D9">species</font></td></tr><tr><td><font color="#99CC00">conductanceScale = PRODUCT OF: conductanceScaling[*]/factor</font></td></tr><tr><td><font color="#99CC00">fopen0 = PRODUCT OF: gates[*]/fcond</font></td></tr><tr><td><font color="#99CC00">fopen = conductanceScale * fopen0</font></td></tr><tr><td><font color="#99CC00">g = conductance * fopen</font></td></tr><tr><td><font color="#666666">Exposures: g (conductance), fopen </font></td></tr></table>>]; ionChannelHH;
|
| 175 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseIonChannel</td></tr><tr><td><font color="#669999">Params: conductance (conductance) </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#B2C0D9">neuroLexId</font></td></tr><tr><td><font color="#666666">Exposures: g (conductance), fopen </font></td></tr></table>>]; baseIonChannel;
|
| 176 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>notes</td></tr></table>>]; notes;
|
| 177 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>gateHHrates</td></tr><tr><td><font color="#FF9966">State vars: q </font></td></tr><tr><td><font color="#99CC00">rateScale = PRODUCT OF: q10Settings[*]/q10</font></td></tr><tr><td><font color="#99CC00">alpha = forwardRate/r</font></td></tr><tr><td><font color="#99CC00">beta = reverseRate/r</font></td></tr><tr><td><font color="#99CC00">fcond = q^instances</font></td></tr><tr><td><font color="#99CC00">inf = alpha/(alpha+beta)</font></td></tr><tr><td><font color="#99CC00">tau = 1/((alpha+beta) * rateScale)</font></td></tr><tr><td><font color="#666633">q' = (inf - q) / tau</font></td></tr><tr><td><font color="#666666">Exposures: alpha (per_time), beta (per_time), tau (time), <br/>inf, rateScale, fcond, <br/>q </font></td></tr></table>>]; gateHHrates;
|
| 178 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>gate</td></tr><tr><td><font color="#666666">Exposures: fcond, q </font></td></tr></table>>]; gate;
|
| 179 |
+
gateHHrates -> gate [len=1.00, arrowhead=onormal]
|
| 180 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseGate</td></tr><tr><td><font color="#669999">Params: instances </font></td></tr><tr><td><font color="#666666">Exposures: fcond, q </font></td></tr></table>>]; baseGate;
|
| 181 |
+
gate -> baseGate [len=1.00, arrowhead=onormal]
|
| 182 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>HHExpLinearRate</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#99CC00">x = (v - midpoint) / scale</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; HHExpLinearRate;
|
| 183 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseHHRate</td></tr><tr><td><font color="#669999">Params: rate (per_time), midpoint (voltage), scale (voltage) </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; baseHHRate;
|
| 184 |
+
HHExpLinearRate -> baseHHRate [len=1.00, arrowhead=onormal]
|
| 185 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseVoltageDepRate</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; baseVoltageDepRate;
|
| 186 |
+
baseHHRate -> baseVoltageDepRate [len=1.00, arrowhead=onormal]
|
| 187 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>HHExpRate</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#99CC00">r = rate * exp((v - midpoint)/scale)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; HHExpRate;
|
| 188 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseHHRate</td></tr><tr><td><font color="#669999">Params: rate (per_time), midpoint (voltage), scale (voltage) </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; baseHHRate;
|
| 189 |
+
HHExpRate -> baseHHRate [len=1.00, arrowhead=onormal]
|
| 190 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseVoltageDepRate</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; baseVoltageDepRate;
|
| 191 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>gateHHrates</td></tr><tr><td><font color="#FF9966">State vars: q </font></td></tr><tr><td><font color="#99CC00">rateScale = PRODUCT OF: q10Settings[*]/q10</font></td></tr><tr><td><font color="#99CC00">alpha = forwardRate/r</font></td></tr><tr><td><font color="#99CC00">beta = reverseRate/r</font></td></tr><tr><td><font color="#99CC00">fcond = q^instances</font></td></tr><tr><td><font color="#99CC00">inf = alpha/(alpha+beta)</font></td></tr><tr><td><font color="#99CC00">tau = 1/((alpha+beta) * rateScale)</font></td></tr><tr><td><font color="#666633">q' = (inf - q) / tau</font></td></tr><tr><td><font color="#666666">Exposures: alpha (per_time), beta (per_time), tau (time), <br/>inf, rateScale, fcond, <br/>q </font></td></tr></table>>]; gateHHrates;
|
| 192 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>gate</td></tr><tr><td><font color="#666666">Exposures: fcond, q </font></td></tr></table>>]; gate;
|
| 193 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseGate</td></tr><tr><td><font color="#669999">Params: instances </font></td></tr><tr><td><font color="#666666">Exposures: fcond, q </font></td></tr></table>>]; baseGate;
|
| 194 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>HHExpRate</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#99CC00">r = rate * exp((v - midpoint)/scale)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; HHExpRate;
|
| 195 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseHHRate</td></tr><tr><td><font color="#669999">Params: rate (per_time), midpoint (voltage), scale (voltage) </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; baseHHRate;
|
| 196 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseVoltageDepRate</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; baseVoltageDepRate;
|
| 197 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>HHSigmoidRate</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#99CC00">r = rate / (1 + exp(0 - (v - midpoint)/scale))</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; HHSigmoidRate;
|
| 198 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseHHRate</td></tr><tr><td><font color="#669999">Params: rate (per_time), midpoint (voltage), scale (voltage) </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; baseHHRate;
|
| 199 |
+
HHSigmoidRate -> baseHHRate [len=1.00, arrowhead=onormal]
|
| 200 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseVoltageDepRate</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; baseVoltageDepRate;
|
| 201 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>channelDensity</td></tr><tr><td><font color="#669999">Params: erev (voltage) </font></td></tr><tr><td><font color="#662211">Consts: vShift = 0 V </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#B2C0D9">segmentGroup</font></td></tr><tr><td><font color="#B2C0D9">ion</font></td></tr><tr><td><font color="#99CC00">channelf = ionChannel/fopen</font></td></tr><tr><td><font color="#99CC00">gDensity = condDensity * channelf</font></td></tr><tr><td><font color="#99CC00">iDensity = gDensity * (erev - v)</font></td></tr><tr><td><font color="#666666">Exposures: gDensity (conductanceDensity), iDensity (currentDensity) </font></td></tr></table>>]; channelDensity;
|
| 202 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseChannelDensityCond</td></tr><tr><td><font color="#669999">Params: condDensity (conductanceDensity) </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: gDensity (conductanceDensity), iDensity (currentDensity) </font></td></tr></table>>]; baseChannelDensityCond;
|
| 203 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseChannelDensity</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: iDensity (currentDensity) </font></td></tr></table>>]; baseChannelDensity;
|
| 204 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>ionChannelHH</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#B2C0D9">species</font></td></tr><tr><td><font color="#99CC00">conductanceScale = PRODUCT OF: conductanceScaling[*]/factor</font></td></tr><tr><td><font color="#99CC00">fopen0 = PRODUCT OF: gates[*]/fcond</font></td></tr><tr><td><font color="#99CC00">fopen = conductanceScale * fopen0</font></td></tr><tr><td><font color="#99CC00">g = conductance * fopen</font></td></tr><tr><td><font color="#666666">Exposures: g (conductance), fopen </font></td></tr></table>>]; ionChannelHH;
|
| 205 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseIonChannel</td></tr><tr><td><font color="#669999">Params: conductance (conductance) </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#B2C0D9">neuroLexId</font></td></tr><tr><td><font color="#666666">Exposures: g (conductance), fopen </font></td></tr></table>>]; baseIonChannel;
|
| 206 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>notes</td></tr></table>>]; notes;
|
| 207 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>gateHHrates</td></tr><tr><td><font color="#FF9966">State vars: q </font></td></tr><tr><td><font color="#99CC00">rateScale = PRODUCT OF: q10Settings[*]/q10</font></td></tr><tr><td><font color="#99CC00">alpha = forwardRate/r</font></td></tr><tr><td><font color="#99CC00">beta = reverseRate/r</font></td></tr><tr><td><font color="#99CC00">fcond = q^instances</font></td></tr><tr><td><font color="#99CC00">inf = alpha/(alpha+beta)</font></td></tr><tr><td><font color="#99CC00">tau = 1/((alpha+beta) * rateScale)</font></td></tr><tr><td><font color="#666633">q' = (inf - q) / tau</font></td></tr><tr><td><font color="#666666">Exposures: alpha (per_time), beta (per_time), tau (time), <br/>inf, rateScale, fcond, <br/>q </font></td></tr></table>>]; gateHHrates;
|
| 208 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>gate</td></tr><tr><td><font color="#666666">Exposures: fcond, q </font></td></tr></table>>]; gate;
|
| 209 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseGate</td></tr><tr><td><font color="#669999">Params: instances </font></td></tr><tr><td><font color="#666666">Exposures: fcond, q </font></td></tr></table>>]; baseGate;
|
| 210 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>HHExpLinearRate</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#99CC00">x = (v - midpoint) / scale</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; HHExpLinearRate;
|
| 211 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseHHRate</td></tr><tr><td><font color="#669999">Params: rate (per_time), midpoint (voltage), scale (voltage) </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; baseHHRate;
|
| 212 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseVoltageDepRate</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; baseVoltageDepRate;
|
| 213 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>HHExpRate</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#99CC00">r = rate * exp((v - midpoint)/scale)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; HHExpRate;
|
| 214 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseHHRate</td></tr><tr><td><font color="#669999">Params: rate (per_time), midpoint (voltage), scale (voltage) </font></td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; baseHHRate;
|
| 215 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>baseVoltageDepRate</td></tr><tr><td><font color="#666699">REQUIRES: v (voltage)</font></td></tr><tr><td><font color="#666666">Exposures: r (per_time) </font></td></tr></table>>]; baseVoltageDepRate;
|
| 216 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>specificCapacitance</td></tr><tr><td><font color="#669999">Params: value (specificCapacitance) </font></td></tr><tr><td><font color="#B2C0D9">segmentGroup</font></td></tr><tr><td><font color="#99CC00">specCap = value</font></td></tr><tr><td><font color="#666666">Exposures: specCap (specificCapacitance) </font></td></tr></table>>]; specificCapacitance;
|
| 217 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>intracellularProperties</td></tr><tr><td><font color="#99CC00">caConc = SUM OF: speciesList[ion='ca']/concentration</font></td></tr><tr><td><font color="#99CC00">caConcExt = SUM OF: speciesList[ion='ca']/extConcentration</font></td></tr><tr><td><font color="#666666">Exposures: caConc (concentration), caConcExt (concentration) </font></td></tr></table>>]; intracellularProperties;
|
| 218 |
+
node [shape=box label=<<table border="0" cellborder="0"><tr><td>resistivity</td></tr><tr><td><font color="#669999">Params: value (resistivity) </font></td></tr><tr><td><font color="#B2C0D9">segmentGroup</font></td></tr></table>>]; resistivity;
|
| 219 |
+
}
|
| 220 |
+
|
| 221 |
+
"hhcell (cell)" -> cell [len=1.00]
|
| 222 |
+
"null (0)" -> notes [len=1.00]
|
| 223 |
+
"morphology (morphology)" -> morphology [len=1.00]
|
| 224 |
+
"soma (id = 0)" -> segment [len=1.00]
|
| 225 |
+
"null (1)" -> proximal [len=1.00]
|
| 226 |
+
"null (2)" -> distal [len=1.00]
|
| 227 |
+
"soma_group (segmentGroup)" -> segmentGroup [len=1.00]
|
| 228 |
+
"null (3)" -> member [len=1.00]
|
| 229 |
+
"bioPhys1 (biophysicalProperties)" -> biophysicalProperties [len=1.00]
|
| 230 |
+
"null (4)" -> membraneProperties [len=1.00]
|
| 231 |
+
"null (5)" -> spikeThresh [len=1.00]
|
| 232 |
+
"null (6)" -> initMembPotential [len=1.00]
|
| 233 |
+
"leak (channelDensity)" -> channelDensity [len=1.00]
|
| 234 |
+
"passiveChan (ionChannelPassive)" -> ionChannelPassive [len=1.00]
|
| 235 |
+
"null (7)" -> notes [len=1.00]
|
| 236 |
+
"naChans (channelDensity)" -> channelDensity [len=1.00]
|
| 237 |
+
"naChan (ionChannelHH)" -> ionChannelHH [len=1.00]
|
| 238 |
+
"null (8)" -> notes [len=1.00]
|
| 239 |
+
"m (gateHHrates)" -> gateHHrates [len=1.00]
|
| 240 |
+
"null (9)" -> HHExpLinearRate [len=1.00]
|
| 241 |
+
"null (10)" -> HHExpRate [len=1.00]
|
| 242 |
+
"h (gateHHrates)" -> gateHHrates [len=1.00]
|
| 243 |
+
"null (11)" -> HHExpRate [len=1.00]
|
| 244 |
+
"null (12)" -> HHSigmoidRate [len=1.00]
|
| 245 |
+
"kChans (channelDensity)" -> channelDensity [len=1.00]
|
| 246 |
+
"kChan (ionChannelHH)" -> ionChannelHH [len=1.00]
|
| 247 |
+
"null (13)" -> notes [len=1.00]
|
| 248 |
+
"n (gateHHrates)" -> gateHHrates [len=1.00]
|
| 249 |
+
"null (14)" -> HHExpLinearRate [len=1.00]
|
| 250 |
+
"null (15)" -> HHExpRate [len=1.00]
|
| 251 |
+
"null (16)" -> specificCapacitance [len=1.00]
|
| 252 |
+
"null (17)" -> intracellularProperties [len=1.00]
|
| 253 |
+
"null (18)" -> resistivity [len=1.00]
|
| 254 |
+
}
|
Tutorial/Source/nmllite/LEMS_SimHHTest.xml
ADDED
|
@@ -0,0 +1,33 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<Lems>
|
| 2 |
+
|
| 3 |
+
<!--
|
| 4 |
+
|
| 5 |
+
This LEMS file has been automatically generated using PyNeuroML v1.1.5 (libNeuroML v0.5.6)
|
| 6 |
+
|
| 7 |
+
-->
|
| 8 |
+
|
| 9 |
+
<!-- Specify which component to run -->
|
| 10 |
+
<Target component="SimHHTest" reportFile="report.SimHHTest.txt"/>
|
| 11 |
+
|
| 12 |
+
<!-- Include core NeuroML2 ComponentType definitions -->
|
| 13 |
+
<Include file="Cells.xml"/>
|
| 14 |
+
<Include file="Networks.xml"/>
|
| 15 |
+
<Include file="Simulation.xml"/>
|
| 16 |
+
|
| 17 |
+
<Include file="PyNN.xml"/>
|
| 18 |
+
<Include file="HHTest.net.nml"/>
|
| 19 |
+
<Include file="/Users/padraig/neuroConstruct/osb/generic/hodgkin_huxley_tutorial/Tutorial/Source/hhcell.cell.nml"/>
|
| 20 |
+
|
| 21 |
+
<Simulation id="SimHHTest" length="50.0ms" step="0.025ms" target="HHTest" seed="5678"> <!-- Note seed: ensures same random numbers used every run -->
|
| 22 |
+
|
| 23 |
+
<Display id="hhPop_0_v" title="Plots of hhPop_0_v" timeScale="1ms" xmin="-5.0" xmax="55.00000000000001" ymin="-80" ymax="40">
|
| 24 |
+
<Line id="hhPop_0__v" quantity="hhPop[0]/v" scale="1mV" color="#d54f33" timeScale="1ms"/>
|
| 25 |
+
</Display>
|
| 26 |
+
|
| 27 |
+
<OutputFile id="hhPop_0_v_dat" fileName="hhPop_0.v.dat">
|
| 28 |
+
<OutputColumn id="hhPop_0__v" quantity="hhPop[0]/v"/>
|
| 29 |
+
</OutputFile>
|
| 30 |
+
|
| 31 |
+
</Simulation>
|
| 32 |
+
|
| 33 |
+
</Lems>
|
Tutorial/Source/nmllite/SimHHTest.json
ADDED
|
@@ -0,0 +1,13 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
{
|
| 2 |
+
"SimHHTest": {
|
| 3 |
+
"version": "NeuroMLlite v0.5.8",
|
| 4 |
+
"network": "HHTest.nmllite.json",
|
| 5 |
+
"duration": 50.0,
|
| 6 |
+
"dt": 0.025,
|
| 7 |
+
"record_variables": {
|
| 8 |
+
"v": {
|
| 9 |
+
"all": "*"
|
| 10 |
+
}
|
| 11 |
+
}
|
| 12 |
+
}
|
| 13 |
+
}
|
Tutorial/Source/nmllite/SimHHTest.yaml
ADDED
|
@@ -0,0 +1,8 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
SimHHTest:
|
| 2 |
+
version: NeuroMLlite v0.5.8
|
| 3 |
+
network: HHTest.nmllite.yaml
|
| 4 |
+
duration: 50.0
|
| 5 |
+
dt: 0.025
|
| 6 |
+
record_variables:
|
| 7 |
+
v:
|
| 8 |
+
all: '*'
|
Tutorial/Source/nmllite/hhPop_0.v.dat
ADDED
|
@@ -0,0 +1,2001 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
+0 -0.065000005
|
| 2 |
+
+2.4999999e-05 -0.064999901
|
| 3 |
+
+4.9999999e-05 -0.064999804
|
| 4 |
+
+7.4999996e-05 -0.064999707
|
| 5 |
+
+9.9999997e-05 -0.064999618
|
| 6 |
+
+0.00012500001 -0.064999521
|
| 7 |
+
+0.00014999999 -0.064999431
|
| 8 |
+
+0.00017499999 -0.064999335
|
| 9 |
+
+0.00019999999 -0.06499923
|
| 10 |
+
+0.000225 -0.064999141
|
| 11 |
+
+0.00025000001 -0.064999051
|
| 12 |
+
+0.000275 -0.064998955
|
| 13 |
+
+0.00029999999 -0.064998865
|
| 14 |
+
+0.000325 -0.064998776
|
| 15 |
+
+0.00034999999 -0.064998694
|
| 16 |
+
+0.000375 -0.064998604
|
| 17 |
+
+0.00039999999 -0.064998515
|
| 18 |
+
+0.00042500001 -0.064998426
|
| 19 |
+
+0.00044999999 -0.064998329
|
| 20 |
+
+0.00047500001 -0.064998247
|
| 21 |
+
+0.00050000002 -0.064998165
|
| 22 |
+
+0.00052499998 -0.06499809
|
| 23 |
+
+0.00055 -0.064998001
|
| 24 |
+
+0.00057500001 -0.064997919
|
| 25 |
+
+0.00059999997 -0.064997844
|
| 26 |
+
+0.00062499999 -0.06499777
|
| 27 |
+
+0.00065 -0.064997688
|
| 28 |
+
+0.00067500002 -0.064997606
|
| 29 |
+
+0.00069999998 -0.064997524
|
| 30 |
+
+0.00072499999 -0.064997442
|
| 31 |
+
+0.00075000001 -0.064997368
|
| 32 |
+
+0.00077500002 -0.064997301
|
| 33 |
+
+0.00079999998 -0.064997226
|
| 34 |
+
+0.000825 -0.064997151
|
| 35 |
+
+0.00085000001 -0.064997077
|
| 36 |
+
+0.00087499997 -0.064996995
|
| 37 |
+
+0.00089999998 -0.064996928
|
| 38 |
+
+0.000925 -0.064996861
|
| 39 |
+
+0.00095000002 -0.064996794
|
| 40 |
+
+0.00097499997 -0.064996719
|
| 41 |
+
+0.001 -0.064996645
|
| 42 |
+
+0.001025 -0.064996578
|
| 43 |
+
+0.00105 -0.064996511
|
| 44 |
+
+0.001075 -0.064996451
|
| 45 |
+
+0.0011 -0.064996377
|
| 46 |
+
+0.001125 -0.06499631
|
| 47 |
+
+0.00115 -0.06499625
|
| 48 |
+
+0.001175 -0.06499619
|
| 49 |
+
+0.0011999999 -0.064996123
|
| 50 |
+
+0.001225 -0.064996049
|
| 51 |
+
+0.00125 -0.064995997
|
| 52 |
+
+0.001275 -0.064995937
|
| 53 |
+
+0.0013 -0.064995877
|
| 54 |
+
+0.001325 -0.064995818
|
| 55 |
+
+0.00135 -0.064995758
|
| 56 |
+
+0.001375 -0.064995699
|
| 57 |
+
+0.0014 -0.064995639
|
| 58 |
+
+0.001425 -0.064995579
|
| 59 |
+
+0.00145 -0.064995527
|
| 60 |
+
+0.0014749999 -0.064995468
|
| 61 |
+
+0.0015 -0.064995416
|
| 62 |
+
+0.001525 -0.064995356
|
| 63 |
+
+0.00155 -0.064995304
|
| 64 |
+
+0.001575 -0.064995252
|
| 65 |
+
+0.0016 -0.064995192
|
| 66 |
+
+0.001625 -0.064995132
|
| 67 |
+
+0.00165 -0.06499508
|
| 68 |
+
+0.0016749999 -0.064995028
|
| 69 |
+
+0.0017 -0.064994983
|
| 70 |
+
+0.001725 -0.064994939
|
| 71 |
+
+0.0017499999 -0.064994887
|
| 72 |
+
+0.001775 -0.064994827
|
| 73 |
+
+0.0018 -0.064994782
|
| 74 |
+
+0.001825 -0.06499473
|
| 75 |
+
+0.00185 -0.064994678
|
| 76 |
+
+0.001875 -0.064994633
|
| 77 |
+
+0.0019 -0.064994588
|
| 78 |
+
+0.001925 -0.064994544
|
| 79 |
+
+0.0019499999 -0.064994499
|
| 80 |
+
+0.0019749999 -0.064994454
|
| 81 |
+
+0.0020000001 -0.06499441
|
| 82 |
+
+0.0020250001 -0.064994372
|
| 83 |
+
+0.00205 -0.064994328
|
| 84 |
+
+0.002075 -0.064994283
|
| 85 |
+
+0.0020999999 -0.064994246
|
| 86 |
+
+0.0021249999 -0.064994194
|
| 87 |
+
+0.0021500001 -0.064994156
|
| 88 |
+
+0.002175 -0.064994127
|
| 89 |
+
+0.0022 -0.064994089
|
| 90 |
+
+0.0022249999 -0.06499406
|
| 91 |
+
+0.0022499999 -0.064994022
|
| 92 |
+
+0.0022750001 -0.064993985
|
| 93 |
+
+0.0023000001 -0.064993955
|
| 94 |
+
+0.002325 -0.064993918
|
| 95 |
+
+0.00235 -0.064993873
|
| 96 |
+
+0.0023749999 -0.064993843
|
| 97 |
+
+0.0023999999 -0.064993806
|
| 98 |
+
+0.0024250001 -0.064993769
|
| 99 |
+
+0.00245 -0.064993739
|
| 100 |
+
+0.002475 -0.064993702
|
| 101 |
+
+0.0024999999 -0.064993672
|
| 102 |
+
+0.0025249999 -0.064993642
|
| 103 |
+
+0.0025500001 -0.064993612
|
| 104 |
+
+0.002575 -0.06499359
|
| 105 |
+
+0.0026 -0.06499356
|
| 106 |
+
+0.002625 -0.064993531
|
| 107 |
+
+0.0026499999 -0.064993508
|
| 108 |
+
+0.0026749999 -0.064993478
|
| 109 |
+
+0.0027000001 -0.064993456
|
| 110 |
+
+0.002725 -0.064993434
|
| 111 |
+
+0.00275 -0.064993404
|
| 112 |
+
+0.0027749999 -0.064993382
|
| 113 |
+
+0.0027999999 -0.064993359
|
| 114 |
+
+0.0028250001 -0.064993337
|
| 115 |
+
+0.00285 -0.064993314
|
| 116 |
+
+0.002875 -0.064993292
|
| 117 |
+
+0.0029 -0.06499327
|
| 118 |
+
+0.0029249999 -0.06499324
|
| 119 |
+
+0.0029499999 -0.064993218
|
| 120 |
+
+0.0029750001 -0.064993195
|
| 121 |
+
+0.003 -0.064993173
|
| 122 |
+
+0.003025 -0.064993151
|
| 123 |
+
+0.0030499999 -0.064993136
|
| 124 |
+
+0.0030749999 -0.064993128
|
| 125 |
+
+0.0031000001 -0.064993113
|
| 126 |
+
+0.003125 -0.064993098
|
| 127 |
+
+0.00315 -0.064993091
|
| 128 |
+
+0.003175 -0.064993076
|
| 129 |
+
+0.0031999999 -0.064993069
|
| 130 |
+
+0.0032249999 -0.064993061
|
| 131 |
+
+0.0032500001 -0.064993046
|
| 132 |
+
+0.003275 -0.064993039
|
| 133 |
+
+0.0033 -0.064993031
|
| 134 |
+
+0.0033249999 -0.064993016
|
| 135 |
+
+0.0033499999 -0.064993009
|
| 136 |
+
+0.0033750001 -0.064993002
|
| 137 |
+
+0.0034 -0.064992994
|
| 138 |
+
+0.003425 -0.064992987
|
| 139 |
+
+0.00345 -0.064992979
|
| 140 |
+
+0.0034749999 -0.064992972
|
| 141 |
+
+0.0034999999 -0.064992964
|
| 142 |
+
+0.0035250001 -0.064992957
|
| 143 |
+
+0.00355 -0.064992949
|
| 144 |
+
+0.003575 -0.064992942
|
| 145 |
+
+0.0035999999 -0.064992942
|
| 146 |
+
+0.0036249999 -0.064992927
|
| 147 |
+
+0.0036500001 -0.06499292
|
| 148 |
+
+0.003675 -0.06499292
|
| 149 |
+
+0.0037 -0.064992912
|
| 150 |
+
+0.003725 -0.064992905
|
| 151 |
+
+0.0037499999 -0.064992905
|
| 152 |
+
+0.0037750001 -0.064992897
|
| 153 |
+
+0.0038000001 -0.064992897
|
| 154 |
+
+0.003825 -0.064992897
|
| 155 |
+
+0.00385 -0.06499289
|
| 156 |
+
+0.0038749999 -0.06499289
|
| 157 |
+
+0.0038999999 -0.06499289
|
| 158 |
+
+0.0039249999 -0.06499289
|
| 159 |
+
+0.0039499998 -0.06499289
|
| 160 |
+
+0.0039749998 -0.06499289
|
| 161 |
+
+0.0040000002 -0.064992897
|
| 162 |
+
+0.0040250001 -0.064992897
|
| 163 |
+
+0.0040500001 -0.064992897
|
| 164 |
+
+0.0040750001 -0.064992905
|
| 165 |
+
+0.0041 -0.064992905
|
| 166 |
+
+0.004125 -0.064992905
|
| 167 |
+
+0.0041499999 -0.064992912
|
| 168 |
+
+0.0041749999 -0.06499292
|
| 169 |
+
+0.0041999999 -0.06499292
|
| 170 |
+
+0.0042249998 -0.064992927
|
| 171 |
+
+0.0042499998 -0.064992942
|
| 172 |
+
+0.0042750002 -0.064992942
|
| 173 |
+
+0.0043000001 -0.064992949
|
| 174 |
+
+0.0043250001 -0.064992957
|
| 175 |
+
+0.0043500001 -0.064992964
|
| 176 |
+
+0.004375 -0.064992972
|
| 177 |
+
+0.0044 -0.064992979
|
| 178 |
+
+0.0044249999 -0.064992987
|
| 179 |
+
+0.0044499999 -0.064993002
|
| 180 |
+
+0.0044749998 -0.064993009
|
| 181 |
+
+0.0044999998 -0.064993009
|
| 182 |
+
+0.0045249998 -0.064993016
|
| 183 |
+
+0.0045500002 -0.064993024
|
| 184 |
+
+0.0045750001 -0.064993024
|
| 185 |
+
+0.0046000001 -0.064993031
|
| 186 |
+
+0.0046250001 -0.064993039
|
| 187 |
+
+0.00465 -0.064993046
|
| 188 |
+
+0.004675 -0.064993054
|
| 189 |
+
+0.0046999999 -0.064993061
|
| 190 |
+
+0.0047249999 -0.064993069
|
| 191 |
+
+0.0047499998 -0.064993076
|
| 192 |
+
+0.0047749998 -0.064993083
|
| 193 |
+
+0.0047999998 -0.064993091
|
| 194 |
+
+0.0048250002 -0.064993098
|
| 195 |
+
+0.0048500001 -0.064993113
|
| 196 |
+
+0.0048750001 -0.064993121
|
| 197 |
+
+0.0049000001 -0.064993128
|
| 198 |
+
+0.004925 -0.064993143
|
| 199 |
+
+0.00495 -0.064993158
|
| 200 |
+
+0.0049749999 -0.064993165
|
| 201 |
+
+0.0049999999 -0.064993173
|
| 202 |
+
+0.0050249998 -0.06622237
|
| 203 |
+
+0.0050499998 -0.067431092
|
| 204 |
+
+0.0050749998 -0.06862092
|
| 205 |
+
+0.0051000002 -0.069793187
|
| 206 |
+
+0.0051250001 -0.070948929
|
| 207 |
+
+0.0051500001 -0.07208889
|
| 208 |
+
+0.0051750001 -0.073213533
|
| 209 |
+
+0.0052 -0.074323088
|
| 210 |
+
+0.005225 -0.07541763
|
| 211 |
+
+0.0052499999 -0.07649713
|
| 212 |
+
+0.0052749999 -0.077561542
|
| 213 |
+
+0.0052999998 -0.078610815
|
| 214 |
+
+0.0053249998 -0.079644941
|
| 215 |
+
+0.0053499998 -0.080663994
|
| 216 |
+
+0.0053750002 -0.081668101
|
| 217 |
+
+0.0054000001 -0.082657449
|
| 218 |
+
+0.0054250001 -0.083632298
|
| 219 |
+
+0.0054500001 -0.084592931
|
| 220 |
+
+0.005475 -0.085539684
|
| 221 |
+
+0.0055 -0.086472914
|
| 222 |
+
+0.0055249999 -0.087392986
|
| 223 |
+
+0.0055499999 -0.08830028
|
| 224 |
+
+0.0055749998 -0.089195177
|
| 225 |
+
+0.0055999998 -0.090078063
|
| 226 |
+
+0.0056249998 -0.090949297
|
| 227 |
+
+0.0056500002 -0.091809258
|
| 228 |
+
+0.0056750001 -0.092658281
|
| 229 |
+
+0.0057000001 -0.093496747
|
| 230 |
+
+0.0057250001 -0.094324984
|
| 231 |
+
+0.00575 -0.095143318
|
| 232 |
+
+0.005775 -0.095952079
|
| 233 |
+
+0.0057999999 -0.096751593
|
| 234 |
+
+0.0058249999 -0.097542152
|
| 235 |
+
+0.0058499998 -0.098324053
|
| 236 |
+
+0.0058749998 -0.099097595
|
| 237 |
+
+0.0058999998 -0.09986303
|
| 238 |
+
+0.0059250002 -0.10062065
|
| 239 |
+
+0.0059500001 -0.10137071
|
| 240 |
+
+0.0059750001 -0.10211346
|
| 241 |
+
+0.0060000001 -0.10284915
|
| 242 |
+
+0.006025 -0.10357801
|
| 243 |
+
+0.00605 -0.10430025
|
| 244 |
+
+0.0060749999 -0.10501612
|
| 245 |
+
+0.0060999999 -0.1057258
|
| 246 |
+
+0.0061249998 -0.10642952
|
| 247 |
+
+0.0061499998 -0.10712745
|
| 248 |
+
+0.0061749998 -0.1078198
|
| 249 |
+
+0.0062000002 -0.10850673
|
| 250 |
+
+0.0062250001 -0.10918843
|
| 251 |
+
+0.0062500001 -0.10986506
|
| 252 |
+
+0.0062750001 -0.11053679
|
| 253 |
+
+0.0063 -0.11120376
|
| 254 |
+
+0.006325 -0.11186612
|
| 255 |
+
+0.0063499999 -0.11252403
|
| 256 |
+
+0.0063749999 -0.1131776
|
| 257 |
+
+0.0063999998 -0.11382698
|
| 258 |
+
+0.0064249998 -0.11447228
|
| 259 |
+
+0.0064499998 -0.11511365
|
| 260 |
+
+0.0064750002 -0.11575115
|
| 261 |
+
+0.0065000001 -0.11638492
|
| 262 |
+
+0.0065250001 -0.11701508
|
| 263 |
+
+0.00655 -0.11764171
|
| 264 |
+
+0.006575 -0.11826489
|
| 265 |
+
+0.0066 -0.11888473
|
| 266 |
+
+0.0066249999 -0.11950132
|
| 267 |
+
+0.0066499999 -0.12011473
|
| 268 |
+
+0.0066749998 -0.12072503
|
| 269 |
+
+0.0066999998 -0.1213323
|
| 270 |
+
+0.0067249998 -0.12193663
|
| 271 |
+
+0.0067500002 -0.12253806
|
| 272 |
+
+0.0067750001 -0.12313666
|
| 273 |
+
+0.0068000001 -0.12373249
|
| 274 |
+
+0.006825 -0.1243256
|
| 275 |
+
+0.00685 -0.12491605
|
| 276 |
+
+0.006875 -0.12550388
|
| 277 |
+
+0.0068999999 -0.12608913
|
| 278 |
+
+0.0069249999 -0.12667185
|
| 279 |
+
+0.0069499998 -0.12725212
|
| 280 |
+
+0.0069749998 -0.12782989
|
| 281 |
+
+0.0069999998 -0.12840527
|
| 282 |
+
+0.0070250002 -0.12897828
|
| 283 |
+
+0.0070500001 -0.12954892
|
| 284 |
+
+0.0070750001 -0.13011724
|
| 285 |
+
+0.0071 -0.13068324
|
| 286 |
+
+0.007125 -0.131247
|
| 287 |
+
+0.00715 -0.13180849
|
| 288 |
+
+0.0071749999 -0.13236775
|
| 289 |
+
+0.0071999999 -0.13292478
|
| 290 |
+
+0.0072249998 -0.13347961
|
| 291 |
+
+0.0072499998 -0.13403225
|
| 292 |
+
+0.0072749997 -0.13458271
|
| 293 |
+
+0.0073000002 -0.13513102
|
| 294 |
+
+0.0073250001 -0.13567716
|
| 295 |
+
+0.0073500001 -0.13622116
|
| 296 |
+
+0.007375 -0.13676301
|
| 297 |
+
+0.0074 -0.13730273
|
| 298 |
+
+0.007425 -0.13784029
|
| 299 |
+
+0.0074499999 -0.13837574
|
| 300 |
+
+0.0074749999 -0.13890904
|
| 301 |
+
+0.0074999998 -0.13944022
|
| 302 |
+
+0.0075249998 -0.13996927
|
| 303 |
+
+0.0075500002 -0.14049619
|
| 304 |
+
+0.0075750002 -0.14102097
|
| 305 |
+
+0.0076000001 -0.14154363
|
| 306 |
+
+0.0076250001 -0.14206414
|
| 307 |
+
+0.00765 -0.14258251
|
| 308 |
+
+0.007675 -0.14309873
|
| 309 |
+
+0.0077 -0.1436128
|
| 310 |
+
+0.0077249999 -0.1441247
|
| 311 |
+
+0.0077499999 -0.14463444
|
| 312 |
+
+0.0077749998 -0.14514199
|
| 313 |
+
+0.0077999998 -0.14564736
|
| 314 |
+
+0.0078250002 -0.14615054
|
| 315 |
+
+0.0078499997 -0.14665154
|
| 316 |
+
+0.0078750001 -0.14715032
|
| 317 |
+
+0.0078999996 -0.14764689
|
| 318 |
+
+0.007925 -0.14814122
|
| 319 |
+
+0.0079499995 -0.14863336
|
| 320 |
+
+0.007975 -0.14912322
|
| 321 |
+
+0.0080000004 -0.14961085
|
| 322 |
+
+0.0080249999 -0.15009621
|
| 323 |
+
+0.0080500003 -0.15057929
|
| 324 |
+
+0.0080749998 -0.1510601
|
| 325 |
+
+0.0081000002 -0.15153864
|
| 326 |
+
+0.0081249997 -0.15201488
|
| 327 |
+
+0.0081500001 -0.15248883
|
| 328 |
+
+0.0081749996 -0.15296045
|
| 329 |
+
+0.0082 -0.15342976
|
| 330 |
+
+0.0082249995 -0.15389673
|
| 331 |
+
+0.00825 -0.15436137
|
| 332 |
+
+0.0082750004 -0.15482363
|
| 333 |
+
+0.0082999999 -0.15528354
|
| 334 |
+
+0.0083250003 -0.1557411
|
| 335 |
+
+0.0083499998 -0.15619628
|
| 336 |
+
+0.0083750002 -0.15664908
|
| 337 |
+
+0.0083999997 -0.15709952
|
| 338 |
+
+0.0084250001 -0.15754753
|
| 339 |
+
+0.0084499996 -0.15799317
|
| 340 |
+
+0.008475 -0.15843639
|
| 341 |
+
+0.0084999995 -0.15887722
|
| 342 |
+
+0.008525 -0.15931563
|
| 343 |
+
+0.0085500004 -0.15975162
|
| 344 |
+
+0.0085749999 -0.16018519
|
| 345 |
+
+0.0086000003 -0.16061632
|
| 346 |
+
+0.0086249998 -0.16104501
|
| 347 |
+
+0.0086500002 -0.16147128
|
| 348 |
+
+0.0086749997 -0.16189508
|
| 349 |
+
+0.0087000001 -0.16231647
|
| 350 |
+
+0.0087249996 -0.1627354
|
| 351 |
+
+0.00875 -0.16315189
|
| 352 |
+
+0.0087749995 -0.16356592
|
| 353 |
+
+0.0088 -0.16397753
|
| 354 |
+
+0.0088250004 -0.16438666
|
| 355 |
+
+0.0088499999 -0.16479336
|
| 356 |
+
+0.0088750003 -0.1651976
|
| 357 |
+
+0.0088999998 -0.16559938
|
| 358 |
+
+0.0089250002 -0.16599871
|
| 359 |
+
+0.0089499997 -0.16639557
|
| 360 |
+
+0.0089750001 -0.16679002
|
| 361 |
+
+0.0089999996 -0.167182
|
| 362 |
+
+0.009025 -0.16757154
|
| 363 |
+
+0.0090499995 -0.16795863
|
| 364 |
+
+0.009075 -0.16834328
|
| 365 |
+
+0.0091000004 -0.16872548
|
| 366 |
+
+0.0091249999 -0.16910525
|
| 367 |
+
+0.0091500003 -0.1694826
|
| 368 |
+
+0.0091749998 -0.16985752
|
| 369 |
+
+0.0092000002 -0.17023003
|
| 370 |
+
+0.0092249997 -0.17060012
|
| 371 |
+
+0.0092500001 -0.17096782
|
| 372 |
+
+0.0092749996 -0.17133307
|
| 373 |
+
+0.0093 -0.17169596
|
| 374 |
+
+0.0093249995 -0.17205645
|
| 375 |
+
+0.0093499999 -0.17241456
|
| 376 |
+
+0.0093750004 -0.17277028
|
| 377 |
+
+0.0093999999 -0.1731236
|
| 378 |
+
+0.0094250003 -0.17347457
|
| 379 |
+
+0.0094499998 -0.17382319
|
| 380 |
+
+0.0094750002 -0.17416945
|
| 381 |
+
+0.0094999997 -0.17451337
|
| 382 |
+
+0.0095250001 -0.17485496
|
| 383 |
+
+0.0095499996 -0.17519423
|
| 384 |
+
+0.009575 -0.17553118
|
| 385 |
+
+0.0095999995 -0.17586581
|
| 386 |
+
+0.0096249999 -0.17619815
|
| 387 |
+
+0.0096500004 -0.1765282
|
| 388 |
+
+0.0096749999 -0.17685598
|
| 389 |
+
+0.0097000003 -0.17718148
|
| 390 |
+
+0.0097249998 -0.17750473
|
| 391 |
+
+0.0097500002 -0.17782573
|
| 392 |
+
+0.0097749997 -0.17814448
|
| 393 |
+
+0.0098000001 -0.178461
|
| 394 |
+
+0.0098249996 -0.17877531
|
| 395 |
+
+0.00985 -0.1790874
|
| 396 |
+
+0.0098749995 -0.17939728
|
| 397 |
+
+0.0098999999 -0.17970501
|
| 398 |
+
+0.0099250004 -0.18001056
|
| 399 |
+
+0.0099499999 -0.18031392
|
| 400 |
+
+0.0099750003 -0.18061511
|
| 401 |
+
+0.0099999998 -0.18091418
|
| 402 |
+
+0.010025 -0.18121111
|
| 403 |
+
+0.01005 -0.18150595
|
| 404 |
+
+0.010075 -0.18179867
|
| 405 |
+
+0.0101 -0.1820893
|
| 406 |
+
+0.010125 -0.18237783
|
| 407 |
+
+0.01015 -0.1826643
|
| 408 |
+
+0.010175 -0.18294871
|
| 409 |
+
+0.0102 -0.18323107
|
| 410 |
+
+0.010225 -0.18351138
|
| 411 |
+
+0.01025 -0.18378967
|
| 412 |
+
+0.010275 -0.18406594
|
| 413 |
+
+0.0103 -0.18434022
|
| 414 |
+
+0.010325 -0.18461253
|
| 415 |
+
+0.01035 -0.18488285
|
| 416 |
+
+0.010375 -0.18515122
|
| 417 |
+
+0.0104 -0.18541762
|
| 418 |
+
+0.010425 -0.18568209
|
| 419 |
+
+0.01045 -0.18594463
|
| 420 |
+
+0.010475 -0.18620525
|
| 421 |
+
+0.0105 -0.18646397
|
| 422 |
+
+0.010525 -0.1867208
|
| 423 |
+
+0.01055 -0.18697576
|
| 424 |
+
+0.010575 -0.18722886
|
| 425 |
+
+0.0106 -0.18748011
|
| 426 |
+
+0.010625 -0.18772951
|
| 427 |
+
+0.01065 -0.18797709
|
| 428 |
+
+0.010675 -0.18822287
|
| 429 |
+
+0.0107 -0.18846682
|
| 430 |
+
+0.010725 -0.18870898
|
| 431 |
+
+0.01075 -0.18894936
|
| 432 |
+
+0.010775 -0.18918797
|
| 433 |
+
+0.0108 -0.18942483
|
| 434 |
+
+0.010825 -0.18965994
|
| 435 |
+
+0.01085 -0.18989332
|
| 436 |
+
+0.010875 -0.19012497
|
| 437 |
+
+0.0109 -0.19035493
|
| 438 |
+
+0.010925 -0.1905832
|
| 439 |
+
+0.01095 -0.19080977
|
| 440 |
+
+0.010975 -0.19103469
|
| 441 |
+
+0.011 -0.19125792
|
| 442 |
+
+0.011025 -0.19147953
|
| 443 |
+
+0.01105 -0.19169949
|
| 444 |
+
+0.011075 -0.19191782
|
| 445 |
+
+0.0111 -0.19213454
|
| 446 |
+
+0.011125 -0.19234966
|
| 447 |
+
+0.01115 -0.19256318
|
| 448 |
+
+0.011175 -0.19277513
|
| 449 |
+
+0.0112 -0.19298549
|
| 450 |
+
+0.011225 -0.1931943
|
| 451 |
+
+0.01125 -0.19340156
|
| 452 |
+
+0.011275 -0.19360729
|
| 453 |
+
+0.0113 -0.19381151
|
| 454 |
+
+0.011325 -0.19401421
|
| 455 |
+
+0.01135 -0.19421537
|
| 456 |
+
+0.011375 -0.19441506
|
| 457 |
+
+0.0114 -0.19461328
|
| 458 |
+
+0.011425 -0.19481003
|
| 459 |
+
+0.01145 -0.19500533
|
| 460 |
+
+0.011475 -0.19519915
|
| 461 |
+
+0.0115 -0.19539155
|
| 462 |
+
+0.011525 -0.19558251
|
| 463 |
+
+0.01155 -0.19577205
|
| 464 |
+
+0.011575 -0.19596019
|
| 465 |
+
+0.0116 -0.19614697
|
| 466 |
+
+0.011625 -0.19633232
|
| 467 |
+
+0.01165 -0.19651629
|
| 468 |
+
+0.011675 -0.19669889
|
| 469 |
+
+0.0117 -0.19688016
|
| 470 |
+
+0.011725 -0.19706008
|
| 471 |
+
+0.01175 -0.19723865
|
| 472 |
+
+0.011775 -0.1974159
|
| 473 |
+
+0.0118 -0.19759184
|
| 474 |
+
+0.011825 -0.19776645
|
| 475 |
+
+0.01185 -0.19793978
|
| 476 |
+
+0.011875 -0.19811183
|
| 477 |
+
+0.0119 -0.19828258
|
| 478 |
+
+0.011925 -0.19845209
|
| 479 |
+
+0.01195 -0.1986203
|
| 480 |
+
+0.011975 -0.19878729
|
| 481 |
+
+0.012 -0.19895302
|
| 482 |
+
+0.012025 -0.19911751
|
| 483 |
+
+0.01205 -0.19928078
|
| 484 |
+
+0.012075 -0.19944285
|
| 485 |
+
+0.0121 -0.19960371
|
| 486 |
+
+0.012125 -0.19976339
|
| 487 |
+
+0.01215 -0.19992189
|
| 488 |
+
+0.012175 -0.20007919
|
| 489 |
+
+0.0122 -0.20023534
|
| 490 |
+
+0.012225 -0.20039029
|
| 491 |
+
+0.01225 -0.20054412
|
| 492 |
+
+0.012275 -0.2006968
|
| 493 |
+
+0.0123 -0.20084834
|
| 494 |
+
+0.012325 -0.20099874
|
| 495 |
+
+0.01235 -0.20114803
|
| 496 |
+
+0.012375 -0.20129621
|
| 497 |
+
+0.0124 -0.2014433
|
| 498 |
+
+0.012425 -0.20158927
|
| 499 |
+
+0.01245 -0.20173417
|
| 500 |
+
+0.012475 -0.20187797
|
| 501 |
+
+0.0125 -0.2020207
|
| 502 |
+
+0.012525 -0.20216237
|
| 503 |
+
+0.01255 -0.20230298
|
| 504 |
+
+0.012575 -0.20244254
|
| 505 |
+
+0.0126 -0.20258108
|
| 506 |
+
+0.012625 -0.20271857
|
| 507 |
+
+0.01265 -0.20285505
|
| 508 |
+
+0.012675 -0.2029905
|
| 509 |
+
+0.0127 -0.20312496
|
| 510 |
+
+0.012725 -0.2032584
|
| 511 |
+
+0.01275 -0.20339087
|
| 512 |
+
+0.012775 -0.20352234
|
| 513 |
+
+0.0128 -0.20365283
|
| 514 |
+
+0.012825 -0.20378235
|
| 515 |
+
+0.01285 -0.2039109
|
| 516 |
+
+0.012875 -0.20403849
|
| 517 |
+
+0.0129 -0.20416515
|
| 518 |
+
+0.012925 -0.20429084
|
| 519 |
+
+0.01295 -0.20441562
|
| 520 |
+
+0.012975 -0.20453945
|
| 521 |
+
+0.013 -0.20466238
|
| 522 |
+
+0.013025 -0.20478439
|
| 523 |
+
+0.01305 -0.20490548
|
| 524 |
+
+0.013075 -0.20502567
|
| 525 |
+
+0.0131 -0.20514497
|
| 526 |
+
+0.013125 -0.20526338
|
| 527 |
+
+0.01315 -0.2053809
|
| 528 |
+
+0.013175 -0.20549755
|
| 529 |
+
+0.0132 -0.20561333
|
| 530 |
+
+0.013225 -0.20572826
|
| 531 |
+
+0.01325 -0.20584235
|
| 532 |
+
+0.013275 -0.20595556
|
| 533 |
+
+0.0133 -0.20606793
|
| 534 |
+
+0.013325 -0.20617948
|
| 535 |
+
+0.01335 -0.2062902
|
| 536 |
+
+0.013375 -0.2064001
|
| 537 |
+
+0.0134 -0.20650916
|
| 538 |
+
+0.013425 -0.20661743
|
| 539 |
+
+0.01345 -0.20672488
|
| 540 |
+
+0.013475 -0.20683151
|
| 541 |
+
+0.0135 -0.20693737
|
| 542 |
+
+0.013525 -0.20704244
|
| 543 |
+
+0.01355 -0.20714673
|
| 544 |
+
+0.013575 -0.20725025
|
| 545 |
+
+0.0136 -0.207353
|
| 546 |
+
+0.013625 -0.20745496
|
| 547 |
+
+0.01365 -0.20755617
|
| 548 |
+
+0.013675 -0.20765664
|
| 549 |
+
+0.0137 -0.20775636
|
| 550 |
+
+0.013725 -0.20785533
|
| 551 |
+
+0.01375 -0.20795356
|
| 552 |
+
+0.013775 -0.20805107
|
| 553 |
+
+0.0138 -0.20814784
|
| 554 |
+
+0.013825 -0.20824389
|
| 555 |
+
+0.01385 -0.20833923
|
| 556 |
+
+0.013875 -0.20843387
|
| 557 |
+
+0.0139 -0.2085278
|
| 558 |
+
+0.013925 -0.20862103
|
| 559 |
+
+0.01395 -0.20871356
|
| 560 |
+
+0.013975 -0.2088054
|
| 561 |
+
+0.014 -0.20889656
|
| 562 |
+
+0.014025 -0.20898704
|
| 563 |
+
+0.01405 -0.20907685
|
| 564 |
+
+0.014075 -0.20916599
|
| 565 |
+
+0.0141 -0.20925447
|
| 566 |
+
+0.014125 -0.2093423
|
| 567 |
+
+0.01415 -0.20942947
|
| 568 |
+
+0.014175 -0.20951599
|
| 569 |
+
+0.0142 -0.20960186
|
| 570 |
+
+0.014225 -0.2096871
|
| 571 |
+
+0.01425 -0.20977169
|
| 572 |
+
+0.014274999 -0.20985566
|
| 573 |
+
+0.0143 -0.209939
|
| 574 |
+
+0.014325 -0.21002172
|
| 575 |
+
+0.01435 -0.21010382
|
| 576 |
+
+0.014375 -0.21018533
|
| 577 |
+
+0.0144 -0.21026622
|
| 578 |
+
+0.014425 -0.21034651
|
| 579 |
+
+0.01445 -0.2104262
|
| 580 |
+
+0.014475 -0.21050531
|
| 581 |
+
+0.0145 -0.21058381
|
| 582 |
+
+0.014525 -0.21066174
|
| 583 |
+
+0.014549999 -0.21073909
|
| 584 |
+
+0.014575 -0.21081585
|
| 585 |
+
+0.0146 -0.21089205
|
| 586 |
+
+0.014625 -0.21096767
|
| 587 |
+
+0.01465 -0.21104275
|
| 588 |
+
+0.014675 -0.21111725
|
| 589 |
+
+0.0147 -0.21119122
|
| 590 |
+
+0.014725 -0.21126461
|
| 591 |
+
+0.01475 -0.21133748
|
| 592 |
+
+0.014775 -0.21140978
|
| 593 |
+
+0.0148 -0.21148156
|
| 594 |
+
+0.014825 -0.2115528
|
| 595 |
+
+0.01485 -0.21162352
|
| 596 |
+
+0.014875 -0.2116937
|
| 597 |
+
+0.0149 -0.21176337
|
| 598 |
+
+0.014925 -0.21183249
|
| 599 |
+
+0.01495 -0.21190113
|
| 600 |
+
+0.014975 -0.21196924
|
| 601 |
+
+0.015 -0.21203686
|
| 602 |
+
+0.015025 -0.21210396
|
| 603 |
+
+0.01505 -0.21217057
|
| 604 |
+
+0.015075 -0.21223669
|
| 605 |
+
+0.0151 -0.2123023
|
| 606 |
+
+0.015125 -0.21236743
|
| 607 |
+
+0.01515 -0.21243207
|
| 608 |
+
+0.015175 -0.21249624
|
| 609 |
+
+0.0152 -0.21255994
|
| 610 |
+
+0.015225 -0.21262315
|
| 611 |
+
+0.01525 -0.2126859
|
| 612 |
+
+0.015275 -0.21274817
|
| 613 |
+
+0.0153 -0.21280998
|
| 614 |
+
+0.015325 -0.21287133
|
| 615 |
+
+0.01535 -0.21293223
|
| 616 |
+
+0.015375 -0.21299267
|
| 617 |
+
+0.0154 -0.21305268
|
| 618 |
+
+0.015425 -0.21311221
|
| 619 |
+
+0.01545 -0.2131713
|
| 620 |
+
+0.015475 -0.21322995
|
| 621 |
+
+0.0155 -0.21328819
|
| 622 |
+
+0.015525 -0.21334597
|
| 623 |
+
+0.01555 -0.21340333
|
| 624 |
+
+0.015575 -0.21346027
|
| 625 |
+
+0.0156 -0.21351679
|
| 626 |
+
+0.015625 -0.21357287
|
| 627 |
+
+0.01565 -0.21362855
|
| 628 |
+
+0.015675001 -0.2136838
|
| 629 |
+
+0.015699999 -0.21373864
|
| 630 |
+
+0.015725 -0.21379308
|
| 631 |
+
+0.01575 -0.21384712
|
| 632 |
+
+0.015775001 -0.21390074
|
| 633 |
+
+0.015799999 -0.21395399
|
| 634 |
+
+0.015825 -0.21400683
|
| 635 |
+
+0.01585 -0.21405928
|
| 636 |
+
+0.015875001 -0.21411133
|
| 637 |
+
+0.015899999 -0.21416299
|
| 638 |
+
+0.015924999 -0.21421428
|
| 639 |
+
+0.01595 -0.21426518
|
| 640 |
+
+0.015975 -0.2143157
|
| 641 |
+
+0.016000001 -0.21436584
|
| 642 |
+
+0.016024999 -0.21441561
|
| 643 |
+
+0.01605 -0.21446501
|
| 644 |
+
+0.016075 -0.21451403
|
| 645 |
+
+0.016100001 -0.2145627
|
| 646 |
+
+0.016124999 -0.21461101
|
| 647 |
+
+0.01615 -0.21465895
|
| 648 |
+
+0.016175 -0.21470654
|
| 649 |
+
+0.0162 -0.21475376
|
| 650 |
+
+0.016225001 -0.21480064
|
| 651 |
+
+0.016249999 -0.21484718
|
| 652 |
+
+0.016275 -0.21489337
|
| 653 |
+
+0.0163 -0.21493921
|
| 654 |
+
+0.016325001 -0.2149847
|
| 655 |
+
+0.016349999 -0.21502987
|
| 656 |
+
+0.016375 -0.2150747
|
| 657 |
+
+0.0164 -0.2151192
|
| 658 |
+
+0.016425001 -0.21516335
|
| 659 |
+
+0.016449999 -0.21520719
|
| 660 |
+
+0.016474999 -0.2152507
|
| 661 |
+
+0.0165 -0.21529388
|
| 662 |
+
+0.016525 -0.21533674
|
| 663 |
+
+0.016550001 -0.21537928
|
| 664 |
+
+0.016574999 -0.2154215
|
| 665 |
+
+0.0166 -0.21546341
|
| 666 |
+
+0.016625 -0.21550502
|
| 667 |
+
+0.016650001 -0.21554631
|
| 668 |
+
+0.016674999 -0.21558729
|
| 669 |
+
+0.0167 -0.21562797
|
| 670 |
+
+0.016725 -0.21566835
|
| 671 |
+
+0.01675 -0.21570842
|
| 672 |
+
+0.016775001 -0.21574819
|
| 673 |
+
+0.016799999 -0.21578766
|
| 674 |
+
+0.016825 -0.21582685
|
| 675 |
+
+0.01685 -0.21586575
|
| 676 |
+
+0.016875001 -0.21590436
|
| 677 |
+
+0.016899999 -0.21594267
|
| 678 |
+
+0.016925 -0.21598069
|
| 679 |
+
+0.01695 -0.21601844
|
| 680 |
+
+0.016975001 -0.2160559
|
| 681 |
+
+0.016999999 -0.21609309
|
| 682 |
+
+0.017024999 -0.21613
|
| 683 |
+
+0.01705 -0.21616663
|
| 684 |
+
+0.017075 -0.216203
|
| 685 |
+
+0.017100001 -0.21623908
|
| 686 |
+
+0.017124999 -0.21627492
|
| 687 |
+
+0.01715 -0.21631047
|
| 688 |
+
+0.017175 -0.21634576
|
| 689 |
+
+0.017200001 -0.21638079
|
| 690 |
+
+0.017224999 -0.21641557
|
| 691 |
+
+0.01725 -0.21645008
|
| 692 |
+
+0.017275 -0.21648434
|
| 693 |
+
+0.0173 -0.21651834
|
| 694 |
+
+0.017325001 -0.21655209
|
| 695 |
+
+0.017349999 -0.21658558
|
| 696 |
+
+0.017375 -0.21661884
|
| 697 |
+
+0.0174 -0.21665184
|
| 698 |
+
+0.017425001 -0.21668458
|
| 699 |
+
+0.017449999 -0.21671708
|
| 700 |
+
+0.017475 -0.21674936
|
| 701 |
+
+0.0175 -0.21678138
|
| 702 |
+
+0.017525 -0.21681316
|
| 703 |
+
+0.017549999 -0.21684472
|
| 704 |
+
+0.017574999 -0.21687603
|
| 705 |
+
+0.0176 -0.21690711
|
| 706 |
+
+0.017625 -0.21693797
|
| 707 |
+
+0.017650001 -0.2169686
|
| 708 |
+
+0.017674999 -0.21699899
|
| 709 |
+
+0.0177 -0.21702915
|
| 710 |
+
+0.017725 -0.21705909
|
| 711 |
+
+0.017750001 -0.21708882
|
| 712 |
+
+0.017774999 -0.21711831
|
| 713 |
+
+0.0178 -0.21714759
|
| 714 |
+
+0.017825 -0.21717666
|
| 715 |
+
+0.01785 -0.21720549
|
| 716 |
+
+0.017875001 -0.21723412
|
| 717 |
+
+0.017899999 -0.21726254
|
| 718 |
+
+0.017925 -0.21729074
|
| 719 |
+
+0.01795 -0.21731876
|
| 720 |
+
+0.017975001 -0.21734653
|
| 721 |
+
+0.017999999 -0.21737413
|
| 722 |
+
+0.018025 -0.2174015
|
| 723 |
+
+0.01805 -0.21742867
|
| 724 |
+
+0.018075 -0.21745564
|
| 725 |
+
+0.018099999 -0.2174824
|
| 726 |
+
+0.018124999 -0.21750897
|
| 727 |
+
+0.01815 -0.21753533
|
| 728 |
+
+0.018175 -0.2175615
|
| 729 |
+
+0.018200001 -0.21758749
|
| 730 |
+
+0.018224999 -0.21761328
|
| 731 |
+
+0.01825 -0.21763887
|
| 732 |
+
+0.018275 -0.21766427
|
| 733 |
+
+0.018300001 -0.2176895
|
| 734 |
+
+0.018324999 -0.21771452
|
| 735 |
+
+0.01835 -0.21773936
|
| 736 |
+
+0.018375 -0.21776402
|
| 737 |
+
+0.0184 -0.21778849
|
| 738 |
+
+0.018425001 -0.21781279
|
| 739 |
+
+0.018449999 -0.21783689
|
| 740 |
+
+0.018475 -0.21786082
|
| 741 |
+
+0.0185 -0.21788457
|
| 742 |
+
+0.018525001 -0.21790816
|
| 743 |
+
+0.018549999 -0.21793155
|
| 744 |
+
+0.018575 -0.21795478
|
| 745 |
+
+0.0186 -0.21797784
|
| 746 |
+
+0.018625 -0.21800072
|
| 747 |
+
+0.018649999 -0.21802343
|
| 748 |
+
+0.018674999 -0.21804596
|
| 749 |
+
+0.0187 -0.21806835
|
| 750 |
+
+0.018725 -0.21809053
|
| 751 |
+
+0.018750001 -0.21811259
|
| 752 |
+
+0.018774999 -0.21813448
|
| 753 |
+
+0.0188 -0.21815619
|
| 754 |
+
+0.018825 -0.21817775
|
| 755 |
+
+0.018850001 -0.21819913
|
| 756 |
+
+0.018874999 -0.21822038
|
| 757 |
+
+0.0189 -0.21824145
|
| 758 |
+
+0.018925 -0.21826237
|
| 759 |
+
+0.01895 -0.21828313
|
| 760 |
+
+0.018975001 -0.21830375
|
| 761 |
+
+0.018999999 -0.21832421
|
| 762 |
+
+0.019025 -0.21834452
|
| 763 |
+
+0.01905 -0.21836469
|
| 764 |
+
+0.019075001 -0.21838468
|
| 765 |
+
+0.019099999 -0.21840453
|
| 766 |
+
+0.019125 -0.21842423
|
| 767 |
+
+0.01915 -0.2184438
|
| 768 |
+
+0.019175 -0.21846321
|
| 769 |
+
+0.019199999 -0.21848248
|
| 770 |
+
+0.019224999 -0.2185016
|
| 771 |
+
+0.01925 -0.21852058
|
| 772 |
+
+0.019275 -0.21853942
|
| 773 |
+
+0.019300001 -0.21855813
|
| 774 |
+
+0.019324999 -0.21857668
|
| 775 |
+
+0.01935 -0.2185951
|
| 776 |
+
+0.019375 -0.21861339
|
| 777 |
+
+0.019400001 -0.21863152
|
| 778 |
+
+0.019424999 -0.21864955
|
| 779 |
+
+0.01945 -0.21866743
|
| 780 |
+
+0.019475 -0.21868518
|
| 781 |
+
+0.0195 -0.21870278
|
| 782 |
+
+0.019525001 -0.21872027
|
| 783 |
+
+0.019549999 -0.21873762
|
| 784 |
+
+0.019575 -0.21875484
|
| 785 |
+
+0.0196 -0.21877193
|
| 786 |
+
+0.019625001 -0.21878891
|
| 787 |
+
+0.019649999 -0.21880575
|
| 788 |
+
+0.019675 -0.21882248
|
| 789 |
+
+0.0197 -0.21883906
|
| 790 |
+
+0.019725 -0.21885553
|
| 791 |
+
+0.019749999 -0.21887186
|
| 792 |
+
+0.019774999 -0.21888809
|
| 793 |
+
+0.0198 -0.21890418
|
| 794 |
+
+0.019825 -0.21892017
|
| 795 |
+
+0.019850001 -0.21893604
|
| 796 |
+
+0.019874999 -0.21895179
|
| 797 |
+
+0.0199 -0.21896742
|
| 798 |
+
+0.019925 -0.21898293
|
| 799 |
+
+0.019950001 -0.21899833
|
| 800 |
+
+0.019974999 -0.21901362
|
| 801 |
+
+0.02 -0.21902879
|
| 802 |
+
+0.020025 -0.21904385
|
| 803 |
+
+0.02005 -0.21905878
|
| 804 |
+
+0.020075001 -0.21907362
|
| 805 |
+
+0.020099999 -0.21908835
|
| 806 |
+
+0.020125 -0.21910296
|
| 807 |
+
+0.02015 -0.21911748
|
| 808 |
+
+0.020175001 -0.21913187
|
| 809 |
+
+0.020199999 -0.21914618
|
| 810 |
+
+0.020225 -0.21916036
|
| 811 |
+
+0.02025 -0.21917444
|
| 812 |
+
+0.020275 -0.21918842
|
| 813 |
+
+0.020299999 -0.21920229
|
| 814 |
+
+0.020324999 -0.21921606
|
| 815 |
+
+0.02035 -0.21922971
|
| 816 |
+
+0.020375 -0.21924329
|
| 817 |
+
+0.020400001 -0.21925674
|
| 818 |
+
+0.020424999 -0.21927011
|
| 819 |
+
+0.02045 -0.21928337
|
| 820 |
+
+0.020475 -0.21929653
|
| 821 |
+
+0.020500001 -0.2193096
|
| 822 |
+
+0.020524999 -0.21932256
|
| 823 |
+
+0.02055 -0.21933542
|
| 824 |
+
+0.020575 -0.21934821
|
| 825 |
+
+0.0206 -0.21936089
|
| 826 |
+
+0.020625001 -0.21937346
|
| 827 |
+
+0.020649999 -0.21938597
|
| 828 |
+
+0.020675 -0.21939836
|
| 829 |
+
+0.0207 -0.21941069
|
| 830 |
+
+0.020725001 -0.21942291
|
| 831 |
+
+0.020749999 -0.21943504
|
| 832 |
+
+0.020775 -0.21944708
|
| 833 |
+
+0.0208 -0.21945903
|
| 834 |
+
+0.020825 -0.21947089
|
| 835 |
+
+0.020849999 -0.21948265
|
| 836 |
+
+0.020874999 -0.21949431
|
| 837 |
+
+0.0209 -0.21950592
|
| 838 |
+
+0.020925 -0.21951742
|
| 839 |
+
+0.020950001 -0.21952884
|
| 840 |
+
+0.020974999 -0.21954018
|
| 841 |
+
+0.021 -0.21955141
|
| 842 |
+
+0.021025 -0.21956259
|
| 843 |
+
+0.021050001 -0.21957366
|
| 844 |
+
+0.021074999 -0.21958466
|
| 845 |
+
+0.0211 -0.2195956
|
| 846 |
+
+0.021125 -0.21960643
|
| 847 |
+
+0.02115 -0.21961719
|
| 848 |
+
+0.021175001 -0.21962786
|
| 849 |
+
+0.021199999 -0.21963847
|
| 850 |
+
+0.021225 -0.21964899
|
| 851 |
+
+0.02125 -0.21965942
|
| 852 |
+
+0.021275001 -0.21966977
|
| 853 |
+
+0.021299999 -0.21968007
|
| 854 |
+
+0.021325 -0.21969028
|
| 855 |
+
+0.02135 -0.21970041
|
| 856 |
+
+0.021375 -0.21971045
|
| 857 |
+
+0.021399999 -0.21972044
|
| 858 |
+
+0.021424999 -0.21973035
|
| 859 |
+
+0.02145 -0.21974018
|
| 860 |
+
+0.021475 -0.21974994
|
| 861 |
+
+0.021500001 -0.21975964
|
| 862 |
+
+0.021524999 -0.21976925
|
| 863 |
+
+0.02155 -0.21977881
|
| 864 |
+
+0.021575 -0.21978828
|
| 865 |
+
+0.021600001 -0.21979769
|
| 866 |
+
+0.021624999 -0.21980701
|
| 867 |
+
+0.02165 -0.21981628
|
| 868 |
+
+0.021675 -0.21982548
|
| 869 |
+
+0.0217 -0.21983461
|
| 870 |
+
+0.021725001 -0.21984367
|
| 871 |
+
+0.021749999 -0.21985266
|
| 872 |
+
+0.021775 -0.21986158
|
| 873 |
+
+0.0218 -0.21987045
|
| 874 |
+
+0.021825001 -0.21987924
|
| 875 |
+
+0.021849999 -0.21988797
|
| 876 |
+
+0.021875 -0.21989663
|
| 877 |
+
+0.0219 -0.21990523
|
| 878 |
+
+0.021925 -0.21991375
|
| 879 |
+
+0.021949999 -0.21992221
|
| 880 |
+
+0.021974999 -0.21993063
|
| 881 |
+
+0.022 -0.21993898
|
| 882 |
+
+0.022025 -0.21994726
|
| 883 |
+
+0.022050001 -0.21995549
|
| 884 |
+
+0.022074999 -0.21996365
|
| 885 |
+
+0.0221 -0.21997175
|
| 886 |
+
+0.022125 -0.21997979
|
| 887 |
+
+0.022150001 -0.21998778
|
| 888 |
+
+0.022174999 -0.21999569
|
| 889 |
+
+0.0222 -0.22000355
|
| 890 |
+
+0.022225 -0.22001135
|
| 891 |
+
+0.02225 -0.22001909
|
| 892 |
+
+0.022275001 -0.22002678
|
| 893 |
+
+0.022299999 -0.22003441
|
| 894 |
+
+0.022325 -0.22004198
|
| 895 |
+
+0.02235 -0.2200495
|
| 896 |
+
+0.022375001 -0.22005695
|
| 897 |
+
+0.022399999 -0.22006436
|
| 898 |
+
+0.022425 -0.22007172
|
| 899 |
+
+0.02245 -0.220079
|
| 900 |
+
+0.022475 -0.22008623
|
| 901 |
+
+0.022499999 -0.22009343
|
| 902 |
+
+0.022524999 -0.22010055
|
| 903 |
+
+0.02255 -0.22010763
|
| 904 |
+
+0.022575 -0.22011465
|
| 905 |
+
+0.022600001 -0.22012162
|
| 906 |
+
+0.022624999 -0.22012855
|
| 907 |
+
+0.02265 -0.22013542
|
| 908 |
+
+0.022675 -0.22014223
|
| 909 |
+
+0.022700001 -0.22014901
|
| 910 |
+
+0.022724999 -0.22015573
|
| 911 |
+
+0.02275 -0.22016239
|
| 912 |
+
+0.022775 -0.22016902
|
| 913 |
+
+0.0228 -0.22017559
|
| 914 |
+
+0.022825001 -0.22018211
|
| 915 |
+
+0.022849999 -0.22018857
|
| 916 |
+
+0.022875 -0.220195
|
| 917 |
+
+0.0229 -0.22020139
|
| 918 |
+
+0.022925001 -0.22020771
|
| 919 |
+
+0.022949999 -0.22021399
|
| 920 |
+
+0.022975 -0.22022024
|
| 921 |
+
+0.023 -0.22022644
|
| 922 |
+
+0.023025 -0.22023259
|
| 923 |
+
+0.023049999 -0.22023869
|
| 924 |
+
+0.023074999 -0.22024475
|
| 925 |
+
+0.0231 -0.22025076
|
| 926 |
+
+0.023125 -0.22025673
|
| 927 |
+
+0.023150001 -0.22026265
|
| 928 |
+
+0.023174999 -0.22026852
|
| 929 |
+
+0.0232 -0.22027434
|
| 930 |
+
+0.023225 -0.22028013
|
| 931 |
+
+0.023250001 -0.22028586
|
| 932 |
+
+0.023274999 -0.22029157
|
| 933 |
+
+0.0233 -0.22029723
|
| 934 |
+
+0.023325 -0.22030285
|
| 935 |
+
+0.02335 -0.22030842
|
| 936 |
+
+0.023375001 -0.22031397
|
| 937 |
+
+0.023399999 -0.22031945
|
| 938 |
+
+0.023425 -0.2203249
|
| 939 |
+
+0.02345 -0.2203303
|
| 940 |
+
+0.023475001 -0.22033568
|
| 941 |
+
+0.023499999 -0.220341
|
| 942 |
+
+0.023525 -0.22034629
|
| 943 |
+
+0.02355 -0.22035155
|
| 944 |
+
+0.023575 -0.22035675
|
| 945 |
+
+0.023599999 -0.22036192
|
| 946 |
+
+0.023624999 -0.22036704
|
| 947 |
+
+0.02365 -0.22037214
|
| 948 |
+
+0.023675 -0.22037719
|
| 949 |
+
+0.023700001 -0.22038221
|
| 950 |
+
+0.023724999 -0.22038721
|
| 951 |
+
+0.02375 -0.22039215
|
| 952 |
+
+0.023775 -0.22039706
|
| 953 |
+
+0.023800001 -0.22040193
|
| 954 |
+
+0.023824999 -0.22040677
|
| 955 |
+
+0.02385 -0.22041157
|
| 956 |
+
+0.023875 -0.22041634
|
| 957 |
+
+0.0239 -0.22042106
|
| 958 |
+
+0.023925001 -0.22042575
|
| 959 |
+
+0.023949999 -0.22043042
|
| 960 |
+
+0.023975 -0.22043504
|
| 961 |
+
+0.024 -0.22043963
|
| 962 |
+
+0.024025001 -0.22044419
|
| 963 |
+
+0.024049999 -0.22044872
|
| 964 |
+
+0.024075 -0.2204532
|
| 965 |
+
+0.0241 -0.22045766
|
| 966 |
+
+0.024125 -0.22046208
|
| 967 |
+
+0.024149999 -0.22046646
|
| 968 |
+
+0.024174999 -0.22047082
|
| 969 |
+
+0.0242 -0.22047514
|
| 970 |
+
+0.024225 -0.22047943
|
| 971 |
+
+0.024250001 -0.22048369
|
| 972 |
+
+0.024274999 -0.22048791
|
| 973 |
+
+0.0243 -0.22049211
|
| 974 |
+
+0.024325 -0.22049627
|
| 975 |
+
+0.024350001 -0.22050041
|
| 976 |
+
+0.024374999 -0.22050451
|
| 977 |
+
+0.0244 -0.22050859
|
| 978 |
+
+0.024425 -0.22051263
|
| 979 |
+
+0.02445 -0.22051664
|
| 980 |
+
+0.024475001 -0.22052063
|
| 981 |
+
+0.024499999 -0.22052458
|
| 982 |
+
+0.024525 -0.2205285
|
| 983 |
+
+0.02455 -0.22053239
|
| 984 |
+
+0.024575001 -0.22053625
|
| 985 |
+
+0.024599999 -0.22054008
|
| 986 |
+
+0.024625 -0.22054389
|
| 987 |
+
+0.02465 -0.22054768
|
| 988 |
+
+0.024675 -0.22055143
|
| 989 |
+
+0.024699999 -0.22055516
|
| 990 |
+
+0.024724999 -0.22055885
|
| 991 |
+
+0.02475 -0.22056252
|
| 992 |
+
+0.024775 -0.22056615
|
| 993 |
+
+0.024800001 -0.22056977
|
| 994 |
+
+0.024824999 -0.22057337
|
| 995 |
+
+0.02485 -0.22057691
|
| 996 |
+
+0.024875 -0.22058046
|
| 997 |
+
+0.024900001 -0.22058396
|
| 998 |
+
+0.024924999 -0.22058745
|
| 999 |
+
+0.02495 -0.22059089
|
| 1000 |
+
+0.024975 -0.22059433
|
| 1001 |
+
+0.025 -0.22059773
|
| 1002 |
+
+0.025025001 -0.2206011
|
| 1003 |
+
+0.025049999 -0.22060445
|
| 1004 |
+
+0.025075 -0.22060779
|
| 1005 |
+
+0.0251 -0.22061108
|
| 1006 |
+
+0.025125001 -0.22061436
|
| 1007 |
+
+0.025149999 -0.22061761
|
| 1008 |
+
+0.025175 -0.22062084
|
| 1009 |
+
+0.0252 -0.22062404
|
| 1010 |
+
+0.025225 -0.22062723
|
| 1011 |
+
+0.025249999 -0.22063039
|
| 1012 |
+
+0.025274999 -0.22063352
|
| 1013 |
+
+0.0253 -0.22063664
|
| 1014 |
+
+0.025325 -0.22063972
|
| 1015 |
+
+0.025350001 -0.22064279
|
| 1016 |
+
+0.025374999 -0.22064583
|
| 1017 |
+
+0.0254 -0.22064884
|
| 1018 |
+
+0.025425 -0.22065184
|
| 1019 |
+
+0.025450001 -0.22065482
|
| 1020 |
+
+0.025474999 -0.22065777
|
| 1021 |
+
+0.0255 -0.2206607
|
| 1022 |
+
+0.025525 -0.22066362
|
| 1023 |
+
+0.02555 -0.2206665
|
| 1024 |
+
+0.025575001 -0.22066937
|
| 1025 |
+
+0.025599999 -0.22067221
|
| 1026 |
+
+0.025625 -0.22067502
|
| 1027 |
+
+0.02565 -0.22067784
|
| 1028 |
+
+0.025675001 -0.22068061
|
| 1029 |
+
+0.025699999 -0.22068338
|
| 1030 |
+
+0.025725 -0.22068612
|
| 1031 |
+
+0.02575 -0.22068883
|
| 1032 |
+
+0.025775 -0.22069153
|
| 1033 |
+
+0.025799999 -0.22069423
|
| 1034 |
+
+0.025824999 -0.22069688
|
| 1035 |
+
+0.02585 -0.22069952
|
| 1036 |
+
+0.025875 -0.22070214
|
| 1037 |
+
+0.025900001 -0.22070473
|
| 1038 |
+
+0.025924999 -0.22070731
|
| 1039 |
+
+0.02595 -0.22070988
|
| 1040 |
+
+0.025975 -0.22071242
|
| 1041 |
+
+0.026000001 -0.22071494
|
| 1042 |
+
+0.026024999 -0.22071745
|
| 1043 |
+
+0.02605 -0.22071993
|
| 1044 |
+
+0.026075 -0.22072241
|
| 1045 |
+
+0.0261 -0.22072487
|
| 1046 |
+
+0.026125001 -0.22072729
|
| 1047 |
+
+0.026149999 -0.22072969
|
| 1048 |
+
+0.026175 -0.22073209
|
| 1049 |
+
+0.0262 -0.22073448
|
| 1050 |
+
+0.026225001 -0.22073685
|
| 1051 |
+
+0.026249999 -0.22073919
|
| 1052 |
+
+0.026275 -0.22074151
|
| 1053 |
+
+0.0263 -0.22074382
|
| 1054 |
+
+0.026325 -0.2207461
|
| 1055 |
+
+0.026349999 -0.22074838
|
| 1056 |
+
+0.026374999 -0.22075063
|
| 1057 |
+
+0.0264 -0.22075288
|
| 1058 |
+
+0.026425 -0.2207551
|
| 1059 |
+
+0.026450001 -0.22075732
|
| 1060 |
+
+0.026474999 -0.22075951
|
| 1061 |
+
+0.0265 -0.22076169
|
| 1062 |
+
+0.026525 -0.22076383
|
| 1063 |
+
+0.026550001 -0.22076598
|
| 1064 |
+
+0.026574999 -0.22076809
|
| 1065 |
+
+0.0266 -0.2207702
|
| 1066 |
+
+0.026625 -0.22077228
|
| 1067 |
+
+0.02665 -0.22077437
|
| 1068 |
+
+0.026675001 -0.22077642
|
| 1069 |
+
+0.026699999 -0.22077847
|
| 1070 |
+
+0.026725 -0.22078049
|
| 1071 |
+
+0.02675 -0.22078252
|
| 1072 |
+
+0.026775001 -0.22078452
|
| 1073 |
+
+0.026799999 -0.2207865
|
| 1074 |
+
+0.026825 -0.22078846
|
| 1075 |
+
+0.02685 -0.22079042
|
| 1076 |
+
+0.026875 -0.22079235
|
| 1077 |
+
+0.026899999 -0.22079428
|
| 1078 |
+
+0.026924999 -0.2207962
|
| 1079 |
+
+0.02695 -0.22079811
|
| 1080 |
+
+0.026975 -0.2208
|
| 1081 |
+
+0.027000001 -0.22080187
|
| 1082 |
+
+0.027024999 -0.22080374
|
| 1083 |
+
+0.02705 -0.22080559
|
| 1084 |
+
+0.027075 -0.22080742
|
| 1085 |
+
+0.027100001 -0.22080924
|
| 1086 |
+
+0.027124999 -0.22081102
|
| 1087 |
+
+0.02715 -0.22081281
|
| 1088 |
+
+0.027175 -0.22081459
|
| 1089 |
+
+0.0272 -0.22081636
|
| 1090 |
+
+0.027225001 -0.22081812
|
| 1091 |
+
+0.027249999 -0.22081985
|
| 1092 |
+
+0.027275 -0.22082157
|
| 1093 |
+
+0.0273 -0.22082329
|
| 1094 |
+
+0.027325001 -0.22082497
|
| 1095 |
+
+0.027349999 -0.22082667
|
| 1096 |
+
+0.027375 -0.22082835
|
| 1097 |
+
+0.0274 -0.22083001
|
| 1098 |
+
+0.027425 -0.22083166
|
| 1099 |
+
+0.027449999 -0.22083329
|
| 1100 |
+
+0.027474999 -0.22083491
|
| 1101 |
+
+0.0275 -0.22083654
|
| 1102 |
+
+0.027525 -0.22083813
|
| 1103 |
+
+0.027550001 -0.22083972
|
| 1104 |
+
+0.027574999 -0.22084129
|
| 1105 |
+
+0.0276 -0.22084285
|
| 1106 |
+
+0.027625 -0.22084442
|
| 1107 |
+
+0.027650001 -0.22084595
|
| 1108 |
+
+0.027674999 -0.22084749
|
| 1109 |
+
+0.0277 -0.22084901
|
| 1110 |
+
+0.027725 -0.22085053
|
| 1111 |
+
+0.02775 -0.22085202
|
| 1112 |
+
+0.027775001 -0.22085349
|
| 1113 |
+
+0.027799999 -0.22085498
|
| 1114 |
+
+0.027825 -0.22085644
|
| 1115 |
+
+0.02785 -0.22085789
|
| 1116 |
+
+0.027875001 -0.22085932
|
| 1117 |
+
+0.027899999 -0.22086076
|
| 1118 |
+
+0.027925 -0.22086218
|
| 1119 |
+
+0.02795 -0.22086358
|
| 1120 |
+
+0.027975 -0.22086498
|
| 1121 |
+
+0.027999999 -0.22086637
|
| 1122 |
+
+0.028024999 -0.22086775
|
| 1123 |
+
+0.02805 -0.22086912
|
| 1124 |
+
+0.028075 -0.22087048
|
| 1125 |
+
+0.028100001 -0.22087182
|
| 1126 |
+
+0.028124999 -0.22087316
|
| 1127 |
+
+0.02815 -0.22087449
|
| 1128 |
+
+0.028175 -0.2208758
|
| 1129 |
+
+0.028200001 -0.22087711
|
| 1130 |
+
+0.028224999 -0.22087841
|
| 1131 |
+
+0.02825 -0.22087969
|
| 1132 |
+
+0.028275 -0.22088097
|
| 1133 |
+
+0.0283 -0.22088224
|
| 1134 |
+
+0.028325001 -0.2208835
|
| 1135 |
+
+0.028349999 -0.22088476
|
| 1136 |
+
+0.028375 -0.22088599
|
| 1137 |
+
+0.0284 -0.22088723
|
| 1138 |
+
+0.028425001 -0.22088845
|
| 1139 |
+
+0.028449999 -0.22088967
|
| 1140 |
+
+0.028475 -0.22089088
|
| 1141 |
+
+0.0285 -0.22089207
|
| 1142 |
+
+0.028525 -0.22089326
|
| 1143 |
+
+0.028549999 -0.22089444
|
| 1144 |
+
+0.028574999 -0.2208956
|
| 1145 |
+
+0.0286 -0.22089677
|
| 1146 |
+
+0.028625 -0.22089791
|
| 1147 |
+
+0.028650001 -0.22089906
|
| 1148 |
+
+0.028674999 -0.22090019
|
| 1149 |
+
+0.0287 -0.22090131
|
| 1150 |
+
+0.028725 -0.22090243
|
| 1151 |
+
+0.028750001 -0.22090355
|
| 1152 |
+
+0.028774999 -0.22090465
|
| 1153 |
+
+0.0288 -0.22090574
|
| 1154 |
+
+0.028825 -0.22090682
|
| 1155 |
+
+0.02885 -0.22090791
|
| 1156 |
+
+0.028875001 -0.22090897
|
| 1157 |
+
+0.028899999 -0.22091004
|
| 1158 |
+
+0.028925 -0.2209111
|
| 1159 |
+
+0.02895 -0.22091214
|
| 1160 |
+
+0.028975001 -0.22091319
|
| 1161 |
+
+0.028999999 -0.22091421
|
| 1162 |
+
+0.029025 -0.22091523
|
| 1163 |
+
+0.02905 -0.22091624
|
| 1164 |
+
+0.029075 -0.22091724
|
| 1165 |
+
+0.029099999 -0.22091825
|
| 1166 |
+
+0.029124999 -0.22091925
|
| 1167 |
+
+0.02915 -0.22092023
|
| 1168 |
+
+0.029175 -0.22092122
|
| 1169 |
+
+0.029200001 -0.22092219
|
| 1170 |
+
+0.029224999 -0.22092316
|
| 1171 |
+
+0.02925 -0.22092411
|
| 1172 |
+
+0.029275 -0.22092506
|
| 1173 |
+
+0.029300001 -0.220926
|
| 1174 |
+
+0.029324999 -0.22092694
|
| 1175 |
+
+0.02935 -0.22092786
|
| 1176 |
+
+0.029375 -0.22092879
|
| 1177 |
+
+0.0294 -0.2209297
|
| 1178 |
+
+0.029425001 -0.22093059
|
| 1179 |
+
+0.029449999 -0.2209315
|
| 1180 |
+
+0.029475 -0.22093239
|
| 1181 |
+
+0.0295 -0.22093329
|
| 1182 |
+
+0.029525001 -0.22093417
|
| 1183 |
+
+0.029549999 -0.22093503
|
| 1184 |
+
+0.029575 -0.22093591
|
| 1185 |
+
+0.0296 -0.22093676
|
| 1186 |
+
+0.029625 -0.22093762
|
| 1187 |
+
+0.029650001 -0.22093847
|
| 1188 |
+
+0.029674999 -0.22093931
|
| 1189 |
+
+0.0297 -0.22094014
|
| 1190 |
+
+0.029725 -0.22094098
|
| 1191 |
+
+0.029750001 -0.2209418
|
| 1192 |
+
+0.029774999 -0.22094262
|
| 1193 |
+
+0.0298 -0.22094344
|
| 1194 |
+
+0.029825 -0.22094424
|
| 1195 |
+
+0.029850001 -0.22094503
|
| 1196 |
+
+0.029874999 -0.22094582
|
| 1197 |
+
+0.029899999 -0.22094662
|
| 1198 |
+
+0.029925 -0.2209474
|
| 1199 |
+
+0.02995 -0.22094817
|
| 1200 |
+
+0.029975001 -0.22094893
|
| 1201 |
+
+0.029999999 -0.22094971
|
| 1202 |
+
+0.030025 -0.22095047
|
| 1203 |
+
+0.03005 -0.21971051
|
| 1204 |
+
+0.030075001 -0.21847981
|
| 1205 |
+
+0.030099999 -0.21725824
|
| 1206 |
+
+0.030125 -0.21604578
|
| 1207 |
+
+0.03015 -0.21484235
|
| 1208 |
+
+0.030175 -0.21364789
|
| 1209 |
+
+0.030200001 -0.21246231
|
| 1210 |
+
+0.030224999 -0.21128555
|
| 1211 |
+
+0.03025 -0.21011755
|
| 1212 |
+
+0.030275 -0.20895825
|
| 1213 |
+
+0.030300001 -0.20780759
|
| 1214 |
+
+0.030324999 -0.20666547
|
| 1215 |
+
+0.03035 -0.20553187
|
| 1216 |
+
+0.030375 -0.20440671
|
| 1217 |
+
+0.030400001 -0.20328991
|
| 1218 |
+
+0.030424999 -0.20218144
|
| 1219 |
+
+0.030449999 -0.20108122
|
| 1220 |
+
+0.030475 -0.19998918
|
| 1221 |
+
+0.0305 -0.19890529
|
| 1222 |
+
+0.030525001 -0.19782944
|
| 1223 |
+
+0.030549999 -0.19676162
|
| 1224 |
+
+0.030575 -0.19570175
|
| 1225 |
+
+0.0306 -0.19464976
|
| 1226 |
+
+0.030625001 -0.1936056
|
| 1227 |
+
+0.030649999 -0.19256923
|
| 1228 |
+
+0.030675 -0.19154055
|
| 1229 |
+
+0.0307 -0.19051954
|
| 1230 |
+
+0.030725 -0.18950613
|
| 1231 |
+
+0.030750001 -0.18850027
|
| 1232 |
+
+0.030774999 -0.18750189
|
| 1233 |
+
+0.0308 -0.18651095
|
| 1234 |
+
+0.030825 -0.18552738
|
| 1235 |
+
+0.030850001 -0.18455113
|
| 1236 |
+
+0.030874999 -0.18358216
|
| 1237 |
+
+0.0309 -0.18262041
|
| 1238 |
+
+0.030925 -0.18166579
|
| 1239 |
+
+0.030950001 -0.1807183
|
| 1240 |
+
+0.030974999 -0.17977786
|
| 1241 |
+
+0.030999999 -0.17884441
|
| 1242 |
+
+0.031025 -0.17791791
|
| 1243 |
+
+0.03105 -0.17699832
|
| 1244 |
+
+0.031075001 -0.17608556
|
| 1245 |
+
+0.031099999 -0.1751796
|
| 1246 |
+
+0.031125 -0.17428039
|
| 1247 |
+
+0.03115 -0.17338787
|
| 1248 |
+
+0.031175001 -0.172502
|
| 1249 |
+
+0.031199999 -0.17162271
|
| 1250 |
+
+0.031225 -0.17074996
|
| 1251 |
+
+0.03125 -0.16988373
|
| 1252 |
+
+0.031275 -0.16902393
|
| 1253 |
+
+0.031300001 -0.16817054
|
| 1254 |
+
+0.031325001 -0.16732351
|
| 1255 |
+
+0.031350002 -0.16648278
|
| 1256 |
+
+0.031374998 -0.16564831
|
| 1257 |
+
+0.031399999 -0.16482005
|
| 1258 |
+
+0.031424999 -0.16399795
|
| 1259 |
+
+0.03145 -0.16318198
|
| 1260 |
+
+0.031475 -0.16237207
|
| 1261 |
+
+0.031500001 -0.16156818
|
| 1262 |
+
+0.031525001 -0.1607703
|
| 1263 |
+
+0.031550001 -0.15997836
|
| 1264 |
+
+0.031575002 -0.15919229
|
| 1265 |
+
+0.031599998 -0.1584121
|
| 1266 |
+
+0.031624999 -0.1576377
|
| 1267 |
+
+0.031649999 -0.15686907
|
| 1268 |
+
+0.031675 -0.15610616
|
| 1269 |
+
+0.0317 -0.15534893
|
| 1270 |
+
+0.031725001 -0.15459734
|
| 1271 |
+
+0.031750001 -0.15385135
|
| 1272 |
+
+0.031775001 -0.15311089
|
| 1273 |
+
+0.031799998 -0.15237597
|
| 1274 |
+
+0.031824999 -0.15164651
|
| 1275 |
+
+0.031849999 -0.15092248
|
| 1276 |
+
+0.031874999 -0.15020384
|
| 1277 |
+
+0.0319 -0.14949055
|
| 1278 |
+
+0.031925 -0.14878257
|
| 1279 |
+
+0.031950001 -0.14807986
|
| 1280 |
+
+0.031975001 -0.14738238
|
| 1281 |
+
+0.032000002 -0.14669009
|
| 1282 |
+
+0.032024998 -0.14600295
|
| 1283 |
+
+0.032049999 -0.14532094
|
| 1284 |
+
+0.032074999 -0.14464399
|
| 1285 |
+
+0.032099999 -0.14397208
|
| 1286 |
+
+0.032125 -0.14330518
|
| 1287 |
+
+0.03215 -0.14264326
|
| 1288 |
+
+0.032175001 -0.14198624
|
| 1289 |
+
+0.032200001 -0.14133413
|
| 1290 |
+
+0.032225002 -0.14068687
|
| 1291 |
+
+0.032249998 -0.14004442
|
| 1292 |
+
+0.032274999 -0.13940676
|
| 1293 |
+
+0.032299999 -0.13877384
|
| 1294 |
+
+0.032325 -0.13814564
|
| 1295 |
+
+0.03235 -0.13752212
|
| 1296 |
+
+0.032375 -0.13690324
|
| 1297 |
+
+0.032400001 -0.13628896
|
| 1298 |
+
+0.032425001 -0.13567924
|
| 1299 |
+
+0.032450002 -0.13507409
|
| 1300 |
+
+0.032474998 -0.13447343
|
| 1301 |
+
+0.032499999 -0.13387723
|
| 1302 |
+
+0.032524999 -0.13328548
|
| 1303 |
+
+0.03255 -0.13269815
|
| 1304 |
+
+0.032575 -0.13211517
|
| 1305 |
+
+0.032600001 -0.13153653
|
| 1306 |
+
+0.032625001 -0.13096221
|
| 1307 |
+
+0.032650001 -0.13039216
|
| 1308 |
+
+0.032675002 -0.12982635
|
| 1309 |
+
+0.032699998 -0.12926474
|
| 1310 |
+
+0.032724999 -0.12870733
|
| 1311 |
+
+0.032749999 -0.12815405
|
| 1312 |
+
+0.032775 -0.12760492
|
| 1313 |
+
+0.0328 -0.12705985
|
| 1314 |
+
+0.032825001 -0.12651886
|
| 1315 |
+
+0.032850001 -0.1259819
|
| 1316 |
+
+0.032875001 -0.12544893
|
| 1317 |
+
+0.032899998 -0.12491993
|
| 1318 |
+
+0.032924999 -0.12439486
|
| 1319 |
+
+0.032949999 -0.12387371
|
| 1320 |
+
+0.032974999 -0.12335644
|
| 1321 |
+
+0.033 -0.12284301
|
| 1322 |
+
+0.033025 -0.12233341
|
| 1323 |
+
+0.033050001 -0.12182761
|
| 1324 |
+
+0.033075001 -0.12132557
|
| 1325 |
+
+0.033100002 -0.12082726
|
| 1326 |
+
+0.033124998 -0.12033267
|
| 1327 |
+
+0.033149999 -0.11984175
|
| 1328 |
+
+0.033174999 -0.11935449
|
| 1329 |
+
+0.033199999 -0.11887085
|
| 1330 |
+
+0.033225 -0.11839082
|
| 1331 |
+
+0.03325 -0.11791436
|
| 1332 |
+
+0.033275001 -0.11744146
|
| 1333 |
+
+0.033300001 -0.11697207
|
| 1334 |
+
+0.033325002 -0.11650617
|
| 1335 |
+
+0.033349998 -0.11604375
|
| 1336 |
+
+0.033374999 -0.11558476
|
| 1337 |
+
+0.033399999 -0.1151292
|
| 1338 |
+
+0.033425 -0.11467701
|
| 1339 |
+
+0.03345 -0.1142282
|
| 1340 |
+
+0.033475 -0.11378273
|
| 1341 |
+
+0.033500001 -0.11334057
|
| 1342 |
+
+0.033525001 -0.11290171
|
| 1343 |
+
+0.033550002 -0.11246612
|
| 1344 |
+
+0.033574998 -0.11203376
|
| 1345 |
+
+0.033599999 -0.11160462
|
| 1346 |
+
+0.033624999 -0.11117869
|
| 1347 |
+
+0.03365 -0.11075592
|
| 1348 |
+
+0.033675 -0.1103363
|
| 1349 |
+
+0.0337 -0.10991979
|
| 1350 |
+
+0.033725001 -0.1095064
|
| 1351 |
+
+0.033750001 -0.10909607
|
| 1352 |
+
+0.033775002 -0.10868881
|
| 1353 |
+
+0.033799998 -0.10828457
|
| 1354 |
+
+0.033824999 -0.10788335
|
| 1355 |
+
+0.033849999 -0.10748512
|
| 1356 |
+
+0.033875 -0.10708984
|
| 1357 |
+
+0.0339 -0.10669751
|
| 1358 |
+
+0.033925001 -0.1063081
|
| 1359 |
+
+0.033950001 -0.10592158
|
| 1360 |
+
+0.033975001 -0.10553794
|
| 1361 |
+
+0.033999998 -0.10515717
|
| 1362 |
+
+0.034024999 -0.10477923
|
| 1363 |
+
+0.034049999 -0.1044041
|
| 1364 |
+
+0.034074999 -0.10403176
|
| 1365 |
+
+0.0341 -0.10366219
|
| 1366 |
+
+0.034125 -0.10329538
|
| 1367 |
+
+0.034150001 -0.10293129
|
| 1368 |
+
+0.034175001 -0.10256992
|
| 1369 |
+
+0.034200002 -0.10221124
|
| 1370 |
+
+0.034224998 -0.10185523
|
| 1371 |
+
+0.034249999 -0.10150187
|
| 1372 |
+
+0.034274999 -0.10115114
|
| 1373 |
+
+0.034299999 -0.10080301
|
| 1374 |
+
+0.034325 -0.10045749
|
| 1375 |
+
+0.03435 -0.10011453
|
| 1376 |
+
+0.034375001 -0.099774122
|
| 1377 |
+
+0.034400001 -0.099436253
|
| 1378 |
+
+0.034425002 -0.099100895
|
| 1379 |
+
+0.034449998 -0.098768033
|
| 1380 |
+
+0.034474999 -0.098437652
|
| 1381 |
+
+0.034499999 -0.09810973
|
| 1382 |
+
+0.034525 -0.097784251
|
| 1383 |
+
+0.03455 -0.097461194
|
| 1384 |
+
+0.034575 -0.097140536
|
| 1385 |
+
+0.034600001 -0.096822269
|
| 1386 |
+
+0.034625001 -0.096506372
|
| 1387 |
+
+0.034650002 -0.096192829
|
| 1388 |
+
+0.034674998 -0.095881619
|
| 1389 |
+
+0.034699999 -0.095572725
|
| 1390 |
+
+0.034724999 -0.095266134
|
| 1391 |
+
+0.03475 -0.094961822
|
| 1392 |
+
+0.034775 -0.094659768
|
| 1393 |
+
+0.0348 -0.094359972
|
| 1394 |
+
+0.034825001 -0.094062403
|
| 1395 |
+
+0.034850001 -0.093767054
|
| 1396 |
+
+0.034875002 -0.093473904
|
| 1397 |
+
+0.034899998 -0.093182936
|
| 1398 |
+
+0.034924999 -0.092894129
|
| 1399 |
+
+0.034949999 -0.092607483
|
| 1400 |
+
+0.034975 -0.092322953
|
| 1401 |
+
+0.035 -0.092040554
|
| 1402 |
+
+0.035025001 -0.091760248
|
| 1403 |
+
+0.035050001 -0.091482036
|
| 1404 |
+
+0.035075001 -0.091205887
|
| 1405 |
+
+0.035099998 -0.090931803
|
| 1406 |
+
+0.035124999 -0.090659752
|
| 1407 |
+
+0.035149999 -0.090389736
|
| 1408 |
+
+0.035174999 -0.090121724
|
| 1409 |
+
+0.0352 -0.089855708
|
| 1410 |
+
+0.035225 -0.089591675
|
| 1411 |
+
+0.035250001 -0.0893296
|
| 1412 |
+
+0.035275001 -0.089069478
|
| 1413 |
+
+0.035300002 -0.088811286
|
| 1414 |
+
+0.035324998 -0.088555016
|
| 1415 |
+
+0.035349999 -0.088300645
|
| 1416 |
+
+0.035374999 -0.088048182
|
| 1417 |
+
+0.035399999 -0.087797597
|
| 1418 |
+
+0.035425 -0.087548867
|
| 1419 |
+
+0.03545 -0.087301999
|
| 1420 |
+
+0.035475001 -0.087056965
|
| 1421 |
+
+0.035500001 -0.086813755
|
| 1422 |
+
+0.035525002 -0.086572357
|
| 1423 |
+
+0.035549998 -0.086332753
|
| 1424 |
+
+0.035574999 -0.086094938
|
| 1425 |
+
+0.035599999 -0.085858874
|
| 1426 |
+
+0.035625 -0.085624576
|
| 1427 |
+
+0.03565 -0.085392021
|
| 1428 |
+
+0.035675 -0.085161187
|
| 1429 |
+
+0.035700001 -0.084932081
|
| 1430 |
+
+0.035725001 -0.084704675
|
| 1431 |
+
+0.035750002 -0.084478967
|
| 1432 |
+
+0.035774998 -0.084254935
|
| 1433 |
+
+0.035799999 -0.084032558
|
| 1434 |
+
+0.035824999 -0.083811842
|
| 1435 |
+
+0.03585 -0.083592772
|
| 1436 |
+
+0.035875 -0.083375335
|
| 1437 |
+
+0.0359 -0.083159491
|
| 1438 |
+
+0.035925001 -0.082945257
|
| 1439 |
+
+0.035950001 -0.082732618
|
| 1440 |
+
+0.035975002 -0.082521565
|
| 1441 |
+
+0.035999998 -0.08231207
|
| 1442 |
+
+0.036024999 -0.082104139
|
| 1443 |
+
+0.036049999 -0.081897743
|
| 1444 |
+
+0.036075 -0.081692874
|
| 1445 |
+
+0.0361 -0.081489533
|
| 1446 |
+
+0.036125001 -0.081287704
|
| 1447 |
+
+0.036150001 -0.081087366
|
| 1448 |
+
+0.036175001 -0.080888517
|
| 1449 |
+
+0.036199998 -0.080691129
|
| 1450 |
+
+0.036224999 -0.080495209
|
| 1451 |
+
+0.036249999 -0.080300733
|
| 1452 |
+
+0.036274999 -0.080107704
|
| 1453 |
+
+0.0363 -0.07991609
|
| 1454 |
+
+0.036325 -0.079725906
|
| 1455 |
+
+0.036350001 -0.079537123
|
| 1456 |
+
+0.036375001 -0.079349726
|
| 1457 |
+
+0.036400001 -0.079163723
|
| 1458 |
+
+0.036424998 -0.078979082
|
| 1459 |
+
+0.036449999 -0.078795806
|
| 1460 |
+
+0.036474999 -0.078613877
|
| 1461 |
+
+0.036499999 -0.07843329
|
| 1462 |
+
+0.036525 -0.078254029
|
| 1463 |
+
+0.03655 -0.078076094
|
| 1464 |
+
+0.036575001 -0.077899449
|
| 1465 |
+
+0.036600001 -0.077724099
|
| 1466 |
+
+0.036625002 -0.077550039
|
| 1467 |
+
+0.036649998 -0.077377245
|
| 1468 |
+
+0.036674999 -0.077205725
|
| 1469 |
+
+0.036699999 -0.077035449
|
| 1470 |
+
+0.036725 -0.076866411
|
| 1471 |
+
+0.03675 -0.076698601
|
| 1472 |
+
+0.036775 -0.076532014
|
| 1473 |
+
+0.036800001 -0.076366641
|
| 1474 |
+
+0.036825001 -0.07620246
|
| 1475 |
+
+0.036850002 -0.076039463
|
| 1476 |
+
+0.036874998 -0.075877644
|
| 1477 |
+
+0.036899999 -0.075716995
|
| 1478 |
+
+0.036924999 -0.075557493
|
| 1479 |
+
+0.03695 -0.075399138
|
| 1480 |
+
+0.036975 -0.075241916
|
| 1481 |
+
+0.037 -0.075085811
|
| 1482 |
+
+0.037025001 -0.074930817
|
| 1483 |
+
+0.037050001 -0.074776925
|
| 1484 |
+
+0.037075002 -0.074624129
|
| 1485 |
+
+0.037099998 -0.074472405
|
| 1486 |
+
+0.037124999 -0.074321747
|
| 1487 |
+
+0.037149999 -0.074172154
|
| 1488 |
+
+0.037175 -0.074023604
|
| 1489 |
+
+0.0372 -0.073876083
|
| 1490 |
+
+0.037225001 -0.073729575
|
| 1491 |
+
+0.037250001 -0.073584087
|
| 1492 |
+
+0.037275001 -0.073439598
|
| 1493 |
+
+0.037299998 -0.0732961
|
| 1494 |
+
+0.037324999 -0.073153578
|
| 1495 |
+
+0.037349999 -0.073012024
|
| 1496 |
+
+0.037374999 -0.072871432
|
| 1497 |
+
+0.0374 -0.072731778
|
| 1498 |
+
+0.037425 -0.072593056
|
| 1499 |
+
+0.037450001 -0.072455257
|
| 1500 |
+
+0.037475001 -0.072318368
|
| 1501 |
+
+0.037500001 -0.072182372
|
| 1502 |
+
+0.037524998 -0.072047263
|
| 1503 |
+
+0.037549999 -0.071913019
|
| 1504 |
+
+0.037574999 -0.071779646
|
| 1505 |
+
+0.037599999 -0.071647115
|
| 1506 |
+
+0.037625 -0.071515426
|
| 1507 |
+
+0.03765 -0.071384557
|
| 1508 |
+
+0.037675001 -0.071254499
|
| 1509 |
+
+0.037700001 -0.071125232
|
| 1510 |
+
+0.037725002 -0.070996754
|
| 1511 |
+
+0.037749998 -0.070869043
|
| 1512 |
+
+0.037774999 -0.070742108
|
| 1513 |
+
+0.037799999 -0.070615903
|
| 1514 |
+
+0.037825 -0.070490442
|
| 1515 |
+
+0.03785 -0.070365705
|
| 1516 |
+
+0.037875 -0.070241667
|
| 1517 |
+
+0.037900001 -0.070118323
|
| 1518 |
+
+0.037925001 -0.069995657
|
| 1519 |
+
+0.037950002 -0.069873661
|
| 1520 |
+
+0.037974998 -0.069752306
|
| 1521 |
+
+0.037999999 -0.069631584
|
| 1522 |
+
+0.038024999 -0.069511503
|
| 1523 |
+
+0.03805 -0.069392018
|
| 1524 |
+
+0.038075 -0.069273129
|
| 1525 |
+
+0.0381 -0.069154814
|
| 1526 |
+
+0.038125001 -0.06903705
|
| 1527 |
+
+0.038150001 -0.068919845
|
| 1528 |
+
+0.038175002 -0.068803161
|
| 1529 |
+
+0.038199998 -0.068686999
|
| 1530 |
+
+0.038224999 -0.068571329
|
| 1531 |
+
+0.038249999 -0.068456143
|
| 1532 |
+
+0.038275 -0.068341419
|
| 1533 |
+
+0.0383 -0.068227142
|
| 1534 |
+
+0.038325001 -0.06811329
|
| 1535 |
+
+0.038350001 -0.067999855
|
| 1536 |
+
+0.038375001 -0.067886822
|
| 1537 |
+
+0.038399998 -0.067774154
|
| 1538 |
+
+0.038424999 -0.067661852
|
| 1539 |
+
+0.038449999 -0.067549884
|
| 1540 |
+
+0.038474999 -0.067438245
|
| 1541 |
+
+0.0385 -0.067326903
|
| 1542 |
+
+0.038525 -0.067215838
|
| 1543 |
+
+0.038550001 -0.06710504
|
| 1544 |
+
+0.038575001 -0.066994473
|
| 1545 |
+
+0.038600001 -0.066884123
|
| 1546 |
+
+0.038624998 -0.066773973
|
| 1547 |
+
+0.038649999 -0.066663995
|
| 1548 |
+
+0.038674999 -0.066554174
|
| 1549 |
+
+0.038699999 -0.066444479
|
| 1550 |
+
+0.038725 -0.066334888
|
| 1551 |
+
+0.03875 -0.06622538
|
| 1552 |
+
+0.038775001 -0.066115923
|
| 1553 |
+
+0.038800001 -0.066006497
|
| 1554 |
+
+0.038825002 -0.065897077
|
| 1555 |
+
+0.038849998 -0.065787621
|
| 1556 |
+
+0.038874999 -0.065678127
|
| 1557 |
+
+0.038899999 -0.065568544
|
| 1558 |
+
+0.038925 -0.065458857
|
| 1559 |
+
+0.03895 -0.065349028
|
| 1560 |
+
+0.038975 -0.065239027
|
| 1561 |
+
+0.039000001 -0.065128818
|
| 1562 |
+
+0.039025001 -0.065018378
|
| 1563 |
+
+0.039050002 -0.064907655
|
| 1564 |
+
+0.039074998 -0.064796634
|
| 1565 |
+
+0.039099999 -0.06468527
|
| 1566 |
+
+0.039124999 -0.064573511
|
| 1567 |
+
+0.03915 -0.064461336
|
| 1568 |
+
+0.039175 -0.064348705
|
| 1569 |
+
+0.0392 -0.064235553
|
| 1570 |
+
+0.039225001 -0.06412185
|
| 1571 |
+
+0.039250001 -0.064007565
|
| 1572 |
+
+0.039275002 -0.063892625
|
| 1573 |
+
+0.039299998 -0.063776992
|
| 1574 |
+
+0.039324999 -0.063660614
|
| 1575 |
+
+0.039349999 -0.063543431
|
| 1576 |
+
+0.039375 -0.063425384
|
| 1577 |
+
+0.0394 -0.063306428
|
| 1578 |
+
+0.039425001 -0.063186489
|
| 1579 |
+
+0.039450001 -0.063065499
|
| 1580 |
+
+0.039475001 -0.062943399
|
| 1581 |
+
+0.039499998 -0.062820099
|
| 1582 |
+
+0.039524999 -0.06269554
|
| 1583 |
+
+0.039549999 -0.062569633
|
| 1584 |
+
+0.039574999 -0.062442306
|
| 1585 |
+
+0.0396 -0.062313452
|
| 1586 |
+
+0.039625 -0.062182985
|
| 1587 |
+
+0.039650001 -0.062050808
|
| 1588 |
+
+0.039675001 -0.06191681
|
| 1589 |
+
+0.039700001 -0.06178087
|
| 1590 |
+
+0.039724998 -0.061642885
|
| 1591 |
+
+0.039749999 -0.061502721
|
| 1592 |
+
+0.039774999 -0.061360247
|
| 1593 |
+
+0.039799999 -0.061215315
|
| 1594 |
+
+0.039825 -0.061067771
|
| 1595 |
+
+0.03985 -0.060917456
|
| 1596 |
+
+0.039875001 -0.06076419
|
| 1597 |
+
+0.039900001 -0.06060778
|
| 1598 |
+
+0.039925002 -0.060448036
|
| 1599 |
+
+0.039949998 -0.060284726
|
| 1600 |
+
+0.039974999 -0.060117625
|
| 1601 |
+
+0.039999999 -0.059946477
|
| 1602 |
+
+0.040025 -0.059771013
|
| 1603 |
+
+0.04005 -0.059590928
|
| 1604 |
+
+0.040075 -0.059405908
|
| 1605 |
+
+0.040100001 -0.059215609
|
| 1606 |
+
+0.040125001 -0.059019644
|
| 1607 |
+
+0.040150002 -0.058817599
|
| 1608 |
+
+0.040174998 -0.058609024
|
| 1609 |
+
+0.040199999 -0.058393423
|
| 1610 |
+
+0.040224999 -0.058170255
|
| 1611 |
+
+0.04025 -0.05793893
|
| 1612 |
+
+0.040275 -0.057698771
|
| 1613 |
+
+0.0403 -0.057449061
|
| 1614 |
+
+0.040325001 -0.057188991
|
| 1615 |
+
+0.040350001 -0.056917664
|
| 1616 |
+
+0.040375002 -0.056634083
|
| 1617 |
+
+0.040399998 -0.056337126
|
| 1618 |
+
+0.040424999 -0.05602555
|
| 1619 |
+
+0.040449999 -0.055697948
|
| 1620 |
+
+0.040475 -0.055352736
|
| 1621 |
+
+0.0405 -0.054988116
|
| 1622 |
+
+0.040525001 -0.054602046
|
| 1623 |
+
+0.040550001 -0.054192219
|
| 1624 |
+
+0.040575001 -0.053755961
|
| 1625 |
+
+0.040599998 -0.053290226
|
| 1626 |
+
+0.040624999 -0.052791484
|
| 1627 |
+
+0.040649999 -0.052255649
|
| 1628 |
+
+0.040674999 -0.051677961
|
| 1629 |
+
+0.0407 -0.05105285
|
| 1630 |
+
+0.040725 -0.050373755
|
| 1631 |
+
+0.040750001 -0.049632929
|
| 1632 |
+
+0.040775001 -0.048821148
|
| 1633 |
+
+0.040800001 -0.047927383
|
| 1634 |
+
+0.040824998 -0.046938371
|
| 1635 |
+
+0.040849999 -0.04583808
|
| 1636 |
+
+0.040874999 -0.044606987
|
| 1637 |
+
+0.040899999 -0.043221265
|
| 1638 |
+
+0.040925 -0.041651629
|
| 1639 |
+
+0.04095 -0.039862003
|
| 1640 |
+
+0.040975001 -0.037807878
|
| 1641 |
+
+0.041000001 -0.035434403
|
| 1642 |
+
+0.041025002 -0.032674391
|
| 1643 |
+
+0.041049998 -0.029446634
|
| 1644 |
+
+0.041074999 -0.025655527
|
| 1645 |
+
+0.041099999 -0.021193881
|
| 1646 |
+
+0.041125 -0.015952518
|
| 1647 |
+
+0.04115 -0.0098423399
|
| 1648 |
+
+0.041175 -0.002836002
|
| 1649 |
+
+0.041200001 +0.0049679494
|
| 1650 |
+
+0.041225001 +0.013273897
|
| 1651 |
+
+0.041250002 +0.021560544
|
| 1652 |
+
+0.041274998 +0.029163491
|
| 1653 |
+
+0.041299999 +0.035482015
|
| 1654 |
+
+0.041324999 +0.040199883
|
| 1655 |
+
+0.04135 +0.04336172
|
| 1656 |
+
+0.041375 +0.045266528
|
| 1657 |
+
+0.0414 +0.046289291
|
| 1658 |
+
+0.041425001 +0.046752207
|
| 1659 |
+
+0.041450001 +0.046879929
|
| 1660 |
+
+0.041475002 +0.046809141
|
| 1661 |
+
+0.041499998 +0.046615694
|
| 1662 |
+
+0.041524999 +0.046339102
|
| 1663 |
+
+0.041549999 +0.045999091
|
| 1664 |
+
+0.041575 +0.045605287
|
| 1665 |
+
+0.0416 +0.045162369
|
| 1666 |
+
+0.041625001 +0.044672653
|
| 1667 |
+
+0.041650001 +0.044137418
|
| 1668 |
+
+0.041675001 +0.043557525
|
| 1669 |
+
+0.041699998 +0.042933721
|
| 1670 |
+
+0.041724999 +0.042266786
|
| 1671 |
+
+0.041749999 +0.041557621
|
| 1672 |
+
+0.041774999 +0.040807243
|
| 1673 |
+
+0.0418 +0.040016841
|
| 1674 |
+
+0.041825 +0.039187718
|
| 1675 |
+
+0.041850001 +0.038321346
|
| 1676 |
+
+0.041875001 +0.037419278
|
| 1677 |
+
+0.041900001 +0.036483217
|
| 1678 |
+
+0.041924998 +0.03551491
|
| 1679 |
+
+0.041949999 +0.034516189
|
| 1680 |
+
+0.041974999 +0.033488944
|
| 1681 |
+
+0.041999999 +0.032435097
|
| 1682 |
+
+0.042025 +0.031356577
|
| 1683 |
+
+0.04205 +0.030255329
|
| 1684 |
+
+0.042075001 +0.029133279
|
| 1685 |
+
+0.042100001 +0.027992329
|
| 1686 |
+
+0.042125002 +0.026834341
|
| 1687 |
+
+0.042149998 +0.025661144
|
| 1688 |
+
+0.042174999 +0.0244745
|
| 1689 |
+
+0.042199999 +0.02327612
|
| 1690 |
+
+0.042225 +0.022067631
|
| 1691 |
+
+0.04225 +0.02085061
|
| 1692 |
+
+0.042275 +0.019626537
|
| 1693 |
+
+0.042300001 +0.018396826
|
| 1694 |
+
+0.042325001 +0.0171628
|
| 1695 |
+
+0.042350002 +0.015925718
|
| 1696 |
+
+0.042374998 +0.014686731
|
| 1697 |
+
+0.042399999 +0.013446923
|
| 1698 |
+
+0.042424999 +0.012207293
|
| 1699 |
+
+0.04245 +0.010968764
|
| 1700 |
+
+0.042475 +0.0097321793
|
| 1701 |
+
+0.0425 +0.0084983036
|
| 1702 |
+
+0.042525001 +0.0072678332
|
| 1703 |
+
+0.042550001 +0.0060413927
|
| 1704 |
+
+0.042575002 +0.0048195389
|
| 1705 |
+
+0.042599998 +0.0036027725
|
| 1706 |
+
+0.042624999 +0.0023915179
|
| 1707 |
+
+0.042649999 +0.0011861591
|
| 1708 |
+
+0.042675 -1.298335e-05
|
| 1709 |
+
+0.0427 -0.00120565
|
| 1710 |
+
+0.042725001 -0.0023916205
|
| 1711 |
+
+0.042750001 -0.003570718
|
| 1712 |
+
+0.042775001 -0.0047428077
|
| 1713 |
+
+0.042799998 -0.0059077991
|
| 1714 |
+
+0.042824998 -0.0070656417
|
| 1715 |
+
+0.042849999 -0.0082163196
|
| 1716 |
+
+0.042874999 -0.0093598608
|
| 1717 |
+
+0.0429 -0.010496311
|
| 1718 |
+
+0.042925 -0.01162576
|
| 1719 |
+
+0.042950001 -0.012748329
|
| 1720 |
+
+0.042975001 -0.01386417
|
| 1721 |
+
+0.043000001 -0.014973469
|
| 1722 |
+
+0.043024998 -0.016076455
|
| 1723 |
+
+0.043049999 -0.017173383
|
| 1724 |
+
+0.043074999 -0.018264556
|
| 1725 |
+
+0.043099999 -0.019350329
|
| 1726 |
+
+0.043125 -0.020431098
|
| 1727 |
+
+0.04315 -0.021507313
|
| 1728 |
+
+0.043175001 -0.022579493
|
| 1729 |
+
+0.043200001 -0.023648219
|
| 1730 |
+
+0.043225002 -0.02471415
|
| 1731 |
+
+0.043249998 -0.025778024
|
| 1732 |
+
+0.043274999 -0.026840694
|
| 1733 |
+
+0.043299999 -0.027903095
|
| 1734 |
+
+0.043325 -0.028966283
|
| 1735 |
+
+0.04335 -0.030031437
|
| 1736 |
+
+0.043375 -0.031099863
|
| 1737 |
+
+0.043400001 -0.032173019
|
| 1738 |
+
+0.043425001 -0.033252504
|
| 1739 |
+
+0.043450002 -0.03434008
|
| 1740 |
+
+0.043474998 -0.035437662
|
| 1741 |
+
+0.043499999 -0.036547322
|
| 1742 |
+
+0.043524999 -0.037671305
|
| 1743 |
+
+0.04355 -0.038811982
|
| 1744 |
+
+0.043575 -0.039971881
|
| 1745 |
+
+0.0436 -0.041153613
|
| 1746 |
+
+0.043625001 -0.04235987
|
| 1747 |
+
+0.043650001 -0.043593343
|
| 1748 |
+
+0.043675002 -0.044856671
|
| 1749 |
+
+0.043699998 -0.046152309
|
| 1750 |
+
+0.043724999 -0.047482416
|
| 1751 |
+
+0.043749999 -0.048848674
|
| 1752 |
+
+0.043775 -0.050252069
|
| 1753 |
+
+0.0438 -0.051692639
|
| 1754 |
+
+0.043825001 -0.053169146
|
| 1755 |
+
+0.043850001 -0.054678775
|
| 1756 |
+
+0.043875001 -0.056216732
|
| 1757 |
+
+0.043899998 -0.057775926
|
| 1758 |
+
+0.043924998 -0.059346717
|
| 1759 |
+
+0.043949999 -0.06091674
|
| 1760 |
+
+0.043974999 -0.062471047
|
| 1761 |
+
+0.044 -0.063992485
|
| 1762 |
+
+0.044025 -0.065462478
|
| 1763 |
+
+0.044050001 -0.066862106
|
| 1764 |
+
+0.044075001 -0.06817358
|
| 1765 |
+
+0.044100001 -0.069381624
|
| 1766 |
+
+0.044124998 -0.070474885
|
| 1767 |
+
+0.044149999 -0.071446829
|
| 1768 |
+
+0.044174999 -0.072296001
|
| 1769 |
+
+0.044199999 -0.073025778
|
| 1770 |
+
+0.044225 -0.073643483
|
| 1771 |
+
+0.04425 -0.074159242
|
| 1772 |
+
+0.044275001 -0.07458479
|
| 1773 |
+
+0.044300001 -0.0749323
|
| 1774 |
+
+0.044325002 -0.075213648
|
| 1775 |
+
+0.044349998 -0.075439751
|
| 1776 |
+
+0.044374999 -0.075620323
|
| 1777 |
+
+0.044399999 -0.075763762
|
| 1778 |
+
+0.044425 -0.0758771
|
| 1779 |
+
+0.04445 -0.075966202
|
| 1780 |
+
+0.044475 -0.076035857
|
| 1781 |
+
+0.044500001 -0.076089941
|
| 1782 |
+
+0.044525001 -0.076131582
|
| 1783 |
+
+0.044550002 -0.07616327
|
| 1784 |
+
+0.044574998 -0.076187022
|
| 1785 |
+
+0.044599999 -0.076204412
|
| 1786 |
+
+0.044624999 -0.076216713
|
| 1787 |
+
+0.04465 -0.076224916
|
| 1788 |
+
+0.044675 -0.076229833
|
| 1789 |
+
+0.0447 -0.076232083
|
| 1790 |
+
+0.044725001 -0.07623218
|
| 1791 |
+
+0.044750001 -0.076230533
|
| 1792 |
+
+0.044775002 -0.076227456
|
| 1793 |
+
+0.044799998 -0.07622321
|
| 1794 |
+
+0.044824999 -0.076218002
|
| 1795 |
+
+0.044849999 -0.076211989
|
| 1796 |
+
+0.044875 -0.076205313
|
| 1797 |
+
+0.0449 -0.076198071
|
| 1798 |
+
+0.044925001 -0.076190352
|
| 1799 |
+
+0.044950001 -0.076182224
|
| 1800 |
+
+0.044975001 -0.076173745
|
| 1801 |
+
+0.044999998 -0.076164961
|
| 1802 |
+
+0.045024998 -0.076155901
|
| 1803 |
+
+0.045049999 -0.076146618
|
| 1804 |
+
+0.045074999 -0.076137111
|
| 1805 |
+
+0.0451 -0.076127417
|
| 1806 |
+
+0.045125 -0.076117538
|
| 1807 |
+
+0.045150001 -0.076107495
|
| 1808 |
+
+0.045175001 -0.076097295
|
| 1809 |
+
+0.045200001 -0.076086946
|
| 1810 |
+
+0.045224998 -0.07607647
|
| 1811 |
+
+0.045249999 -0.076065853
|
| 1812 |
+
+0.045274999 -0.07605511
|
| 1813 |
+
+0.045299999 -0.076044247
|
| 1814 |
+
+0.045325 -0.076033249
|
| 1815 |
+
+0.04535 -0.076022141
|
| 1816 |
+
+0.045375001 -0.076010928
|
| 1817 |
+
+0.045400001 -0.075999595
|
| 1818 |
+
+0.045425002 -0.075988136
|
| 1819 |
+
+0.045449998 -0.075976573
|
| 1820 |
+
+0.045474999 -0.075964883
|
| 1821 |
+
+0.045499999 -0.075953089
|
| 1822 |
+
+0.045524999 -0.075941198
|
| 1823 |
+
+0.04555 -0.07592918
|
| 1824 |
+
+0.045575 -0.075917058
|
| 1825 |
+
+0.045600001 -0.075904824
|
| 1826 |
+
+0.045625001 -0.075892493
|
| 1827 |
+
+0.045650002 -0.075880028
|
| 1828 |
+
+0.045674998 -0.075867474
|
| 1829 |
+
+0.045699999 -0.075854801
|
| 1830 |
+
+0.045724999 -0.075842023
|
| 1831 |
+
+0.04575 -0.075829126
|
| 1832 |
+
+0.045775 -0.075816125
|
| 1833 |
+
+0.0458 -0.075803012
|
| 1834 |
+
+0.045825001 -0.075789787
|
| 1835 |
+
+0.045850001 -0.07577645
|
| 1836 |
+
+0.045875002 -0.075762995
|
| 1837 |
+
+0.045899998 -0.075749435
|
| 1838 |
+
+0.045924999 -0.075735755
|
| 1839 |
+
+0.045949999 -0.075721964
|
| 1840 |
+
+0.045975 -0.075708061
|
| 1841 |
+
+0.046 -0.075694047
|
| 1842 |
+
+0.046025001 -0.075679921
|
| 1843 |
+
+0.046050001 -0.075665675
|
| 1844 |
+
+0.046075001 -0.075651318
|
| 1845 |
+
+0.046099998 -0.075636849
|
| 1846 |
+
+0.046124998 -0.075622275
|
| 1847 |
+
+0.046149999 -0.075607575
|
| 1848 |
+
+0.046174999 -0.075592771
|
| 1849 |
+
+0.0462 -0.07557784
|
| 1850 |
+
+0.046225 -0.07556279
|
| 1851 |
+
+0.046250001 -0.075547636
|
| 1852 |
+
+0.046275001 -0.075532362
|
| 1853 |
+
+0.046300001 -0.075516976
|
| 1854 |
+
+0.046324998 -0.075501472
|
| 1855 |
+
+0.046349999 -0.07548584
|
| 1856 |
+
+0.046374999 -0.075470105
|
| 1857 |
+
+0.046399999 -0.07545425
|
| 1858 |
+
+0.046425 -0.075438276
|
| 1859 |
+
+0.04645 -0.075422183
|
| 1860 |
+
+0.046475001 -0.075405985
|
| 1861 |
+
+0.046500001 -0.075389668
|
| 1862 |
+
+0.046525002 -0.075373232
|
| 1863 |
+
+0.046549998 -0.075356677
|
| 1864 |
+
+0.046574999 -0.07534001
|
| 1865 |
+
+0.046599999 -0.075323217
|
| 1866 |
+
+0.046624999 -0.075306311
|
| 1867 |
+
+0.04665 -0.075289287
|
| 1868 |
+
+0.046675 -0.07527215
|
| 1869 |
+
+0.046700001 -0.075254902
|
| 1870 |
+
+0.046725001 -0.075237527
|
| 1871 |
+
+0.046750002 -0.075220034
|
| 1872 |
+
+0.046774998 -0.075202428
|
| 1873 |
+
+0.046799999 -0.075184703
|
| 1874 |
+
+0.046824999 -0.075166859
|
| 1875 |
+
+0.04685 -0.075148895
|
| 1876 |
+
+0.046875 -0.07513082
|
| 1877 |
+
+0.0469 -0.075112626
|
| 1878 |
+
+0.046925001 -0.075094312
|
| 1879 |
+
+0.046950001 -0.075075887
|
| 1880 |
+
+0.046975002 -0.075057335
|
| 1881 |
+
+0.046999998 -0.075038671
|
| 1882 |
+
+0.047024999 -0.075019896
|
| 1883 |
+
+0.047049999 -0.075001001
|
| 1884 |
+
+0.047075 -0.074981987
|
| 1885 |
+
+0.0471 -0.074962862
|
| 1886 |
+
+0.047125001 -0.074943624
|
| 1887 |
+
+0.047150001 -0.074924268
|
| 1888 |
+
+0.047175001 -0.074904799
|
| 1889 |
+
+0.047199998 -0.074885212
|
| 1890 |
+
+0.047224998 -0.074865513
|
| 1891 |
+
+0.047249999 -0.074845701
|
| 1892 |
+
+0.047274999 -0.074825771
|
| 1893 |
+
+0.0473 -0.074805729
|
| 1894 |
+
+0.047325 -0.074785575
|
| 1895 |
+
+0.047350001 -0.074765302
|
| 1896 |
+
+0.047375001 -0.074744917
|
| 1897 |
+
+0.047400001 -0.074724421
|
| 1898 |
+
+0.047424998 -0.074703813
|
| 1899 |
+
+0.047449999 -0.074683093
|
| 1900 |
+
+0.047474999 -0.074662261
|
| 1901 |
+
+0.047499999 -0.074641317
|
| 1902 |
+
+0.047525 -0.074620269
|
| 1903 |
+
+0.04755 -0.074599095
|
| 1904 |
+
+0.047575001 -0.074577816
|
| 1905 |
+
+0.047600001 -0.07455644
|
| 1906 |
+
+0.047625002 -0.074534945
|
| 1907 |
+
+0.047649998 -0.074513346
|
| 1908 |
+
+0.047674999 -0.074491635
|
| 1909 |
+
+0.047699999 -0.074469812
|
| 1910 |
+
+0.047724999 -0.074447885
|
| 1911 |
+
+0.04775 -0.074425861
|
| 1912 |
+
+0.047775 -0.074403726
|
| 1913 |
+
+0.047800001 -0.074381478
|
| 1914 |
+
+0.047825001 -0.074359134
|
| 1915 |
+
+0.047850002 -0.074336678
|
| 1916 |
+
+0.047874998 -0.074314117
|
| 1917 |
+
+0.047899999 -0.07429146
|
| 1918 |
+
+0.047924999 -0.074268699
|
| 1919 |
+
+0.04795 -0.074245825
|
| 1920 |
+
+0.047975 -0.074222848
|
| 1921 |
+
+0.048 -0.074199781
|
| 1922 |
+
+0.048025001 -0.07417661
|
| 1923 |
+
+0.048050001 -0.074153334
|
| 1924 |
+
+0.048075002 -0.074129961
|
| 1925 |
+
+0.048099998 -0.074106485
|
| 1926 |
+
+0.048124999 -0.074082911
|
| 1927 |
+
+0.048149999 -0.074059255
|
| 1928 |
+
+0.048175 -0.074035488
|
| 1929 |
+
+0.0482 -0.074011631
|
| 1930 |
+
+0.048225001 -0.073987685
|
| 1931 |
+
+0.048250001 -0.073963635
|
| 1932 |
+
+0.048275001 -0.073939495
|
| 1933 |
+
+0.048299998 -0.073915258
|
| 1934 |
+
+0.048324998 -0.073890917
|
| 1935 |
+
+0.048349999 -0.073866494
|
| 1936 |
+
+0.048374999 -0.073841982
|
| 1937 |
+
+0.0484 -0.073817387
|
| 1938 |
+
+0.048425 -0.073792696
|
| 1939 |
+
+0.048450001 -0.073767908
|
| 1940 |
+
+0.048475001 -0.073743045
|
| 1941 |
+
+0.048500001 -0.073718101
|
| 1942 |
+
+0.048524998 -0.073693059
|
| 1943 |
+
+0.048549999 -0.073667936
|
| 1944 |
+
+0.048574999 -0.073642723
|
| 1945 |
+
+0.048599999 -0.073617443
|
| 1946 |
+
+0.048625 -0.073592074
|
| 1947 |
+
+0.04865 -0.073566616
|
| 1948 |
+
+0.048675001 -0.073541075
|
| 1949 |
+
+0.048700001 -0.073515452
|
| 1950 |
+
+0.048725002 -0.073489748
|
| 1951 |
+
+0.048749998 -0.073463969
|
| 1952 |
+
+0.048774999 -0.073438115
|
| 1953 |
+
+0.048799999 -0.07341218
|
| 1954 |
+
+0.048824999 -0.07338617
|
| 1955 |
+
+0.04885 -0.073360093
|
| 1956 |
+
+0.048875 -0.073333941
|
| 1957 |
+
+0.048900001 -0.073307715
|
| 1958 |
+
+0.048925001 -0.073281415
|
| 1959 |
+
+0.048950002 -0.07325504
|
| 1960 |
+
+0.048974998 -0.073228598
|
| 1961 |
+
+0.048999999 -0.073202074
|
| 1962 |
+
+0.049024999 -0.073175497
|
| 1963 |
+
+0.04905 -0.073148847
|
| 1964 |
+
+0.049075 -0.073122129
|
| 1965 |
+
+0.0491 -0.073095351
|
| 1966 |
+
+0.049125001 -0.0730685
|
| 1967 |
+
+0.049150001 -0.073041588
|
| 1968 |
+
+0.049175002 -0.073014624
|
| 1969 |
+
+0.049199998 -0.072987594
|
| 1970 |
+
+0.049224999 -0.072960496
|
| 1971 |
+
+0.049249999 -0.072933346
|
| 1972 |
+
+0.049275 -0.072906129
|
| 1973 |
+
+0.0493 -0.072878867
|
| 1974 |
+
+0.049325 -0.072851539
|
| 1975 |
+
+0.049350001 -0.07282415
|
| 1976 |
+
+0.049375001 -0.07279671
|
| 1977 |
+
+0.049399998 -0.072769217
|
| 1978 |
+
+0.049424998 -0.072741672
|
| 1979 |
+
+0.049449999 -0.072714075
|
| 1980 |
+
+0.049474999 -0.072686426
|
| 1981 |
+
+0.0495 -0.072658733
|
| 1982 |
+
+0.049525 -0.072630987
|
| 1983 |
+
+0.049550001 -0.072603188
|
| 1984 |
+
+0.049575001 -0.072575353
|
| 1985 |
+
+0.049600001 -0.072547466
|
| 1986 |
+
+0.049624998 -0.072519533
|
| 1987 |
+
+0.049649999 -0.072491564
|
| 1988 |
+
+0.049674999 -0.072463557
|
| 1989 |
+
+0.049699999 -0.072435506
|
| 1990 |
+
+0.049725 -0.07240741
|
| 1991 |
+
+0.04975 -0.072379284
|
| 1992 |
+
+0.049775001 -0.072351113
|
| 1993 |
+
+0.049800001 -0.072322913
|
| 1994 |
+
+0.049825002 -0.072294667
|
| 1995 |
+
+0.049849998 -0.072266392
|
| 1996 |
+
+0.049874999 -0.072238073
|
| 1997 |
+
+0.049899999 -0.072209731
|
| 1998 |
+
+0.049924999 -0.072181351
|
| 1999 |
+
+0.04995 -0.07215295
|
| 2000 |
+
+0.049975 -0.072124511
|
| 2001 |
+
+0.050000001 -0.072096042
|
Tutorial/Source/nmllite/hhcell.cell.nml
ADDED
|
@@ -0,0 +1,51 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<?xml version="1.0" encoding="UTF-8"?>
|
| 2 |
+
|
| 3 |
+
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2"
|
| 4 |
+
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
|
| 5 |
+
xsi:schemaLocation="http://www.neuroml.org/schema/neuroml2 https://raw.githubusercontent.com/NeuroML/NeuroML2/master/Schemas/NeuroML2/NeuroML_v2.2.xsd"
|
| 6 |
+
id="hhcell">
|
| 7 |
+
|
| 8 |
+
<include href="passiveChan.channel.nml"/> <!-- Include the channel definitions -->
|
| 9 |
+
<include href="naChan.channel.nml"/>
|
| 10 |
+
<include href="kChan.channel.nml"/>
|
| 11 |
+
|
| 12 |
+
<cell id="hhcell">
|
| 13 |
+
|
| 14 |
+
<notes>Conductance based cell model NeuroML2 format: standard Hodgkin Huxley model cell with Na, K and passive conductances</notes>
|
| 15 |
+
|
| 16 |
+
<morphology id="morphology">
|
| 17 |
+
<segment id="0" name="soma">
|
| 18 |
+
<proximal x="0" y="0" z="0" diameter="17.841242"/> <!--Gives a convenient surface area of 1000.0 um^2-->
|
| 19 |
+
<distal x="0" y="0" z="0" diameter="17.841242"/>
|
| 20 |
+
</segment>
|
| 21 |
+
|
| 22 |
+
<segmentGroup id="soma_group">
|
| 23 |
+
<member segment="0"/>
|
| 24 |
+
</segmentGroup>
|
| 25 |
+
|
| 26 |
+
</morphology>
|
| 27 |
+
|
| 28 |
+
<biophysicalProperties id="bioPhys1">
|
| 29 |
+
|
| 30 |
+
<membraneProperties>
|
| 31 |
+
|
| 32 |
+
<channelDensity id="leak" ionChannel="passiveChan" condDensity="0.3 mS_per_cm2" erev="-54.387mV" ion="non_specific"/>
|
| 33 |
+
<channelDensity id="naChans" ionChannel="naChan" condDensity="120.0 mS_per_cm2" erev="50.0 mV" ion="na"/>
|
| 34 |
+
<channelDensity id="kChans" ionChannel="kChan" condDensity="36 mS_per_cm2" erev="-77mV" ion="k"/>
|
| 35 |
+
|
| 36 |
+
<spikeThresh value="-20mV"/>
|
| 37 |
+
<specificCapacitance value="1.0 uF_per_cm2"/>
|
| 38 |
+
<initMembPotential value="-65mV"/>
|
| 39 |
+
|
| 40 |
+
</membraneProperties>
|
| 41 |
+
|
| 42 |
+
<intracellularProperties>
|
| 43 |
+
<resistivity value="0.03 kohm_cm"/> <!-- Note: not used in single compartment simulations -->
|
| 44 |
+
</intracellularProperties>
|
| 45 |
+
|
| 46 |
+
</biophysicalProperties>
|
| 47 |
+
|
| 48 |
+
</cell>
|
| 49 |
+
|
| 50 |
+
|
| 51 |
+
</neuroml>
|
Tutorial/Source/nmllite/kChan.channel.nml
ADDED
|
@@ -0,0 +1,22 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<?xml version="1.0" encoding="UTF-8"?>
|
| 2 |
+
|
| 3 |
+
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2"
|
| 4 |
+
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
|
| 5 |
+
xsi:schemaLocation="http://www.neuroml.org/schema/neuroml2 https://raw.githubusercontent.com/NeuroML/NeuroML2/master/Schemas/NeuroML2/NeuroML_v2beta3.xsd"
|
| 6 |
+
id="kChan">
|
| 7 |
+
|
| 8 |
+
|
| 9 |
+
<ionChannelHH id="kChan" conductance="10pS" species="k">
|
| 10 |
+
|
| 11 |
+
<notes>Single ion channel in NeuroML2 format: standard Potassium channel from the Hodgkin Huxley model</notes>
|
| 12 |
+
|
| 13 |
+
<gateHHrates id="n" instances="4">
|
| 14 |
+
<forwardRate type="HHExpLinearRate" rate="0.1per_ms" midpoint="-55mV" scale="10mV"/>
|
| 15 |
+
<reverseRate type="HHExpRate" rate="0.125per_ms" midpoint="-65mV" scale="-80mV"/>
|
| 16 |
+
</gateHHrates>
|
| 17 |
+
|
| 18 |
+
</ionChannelHH>
|
| 19 |
+
|
| 20 |
+
|
| 21 |
+
</neuroml>
|
| 22 |
+
|
Tutorial/Source/nmllite/naChan.channel.nml
ADDED
|
@@ -0,0 +1,27 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<?xml version="1.0" encoding="UTF-8"?>
|
| 2 |
+
|
| 3 |
+
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2"
|
| 4 |
+
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
|
| 5 |
+
xsi:schemaLocation="http://www.neuroml.org/schema/neuroml2 https://raw.githubusercontent.com/NeuroML/NeuroML2/master/Schemas/NeuroML2/NeuroML_v2beta3.xsd"
|
| 6 |
+
id="naChan">
|
| 7 |
+
|
| 8 |
+
|
| 9 |
+
<ionChannelHH id="naChan" conductance="10pS" species="na">
|
| 10 |
+
|
| 11 |
+
<notes>Single ion channel in NeuroML2 format: standard Sodium channel from the Hodgkin Huxley model</notes>
|
| 12 |
+
|
| 13 |
+
<gateHHrates id="m" instances="3">
|
| 14 |
+
<forwardRate type="HHExpLinearRate" rate="1per_ms" midpoint="-40mV" scale="10mV"/>
|
| 15 |
+
<reverseRate type="HHExpRate" rate="4per_ms" midpoint="-65mV" scale="-18mV"/>
|
| 16 |
+
</gateHHrates>
|
| 17 |
+
|
| 18 |
+
<gateHHrates id="h" instances="1">
|
| 19 |
+
<forwardRate type="HHExpRate" rate="0.07per_ms" midpoint="-65mV" scale="-20mV"/>
|
| 20 |
+
<reverseRate type="HHSigmoidRate" rate="1per_ms" midpoint="-35mV" scale="10mV"/>
|
| 21 |
+
</gateHHrates>
|
| 22 |
+
|
| 23 |
+
</ionChannelHH>
|
| 24 |
+
|
| 25 |
+
|
| 26 |
+
</neuroml>
|
| 27 |
+
|
Tutorial/Source/nmllite/passiveChan.channel.nml
ADDED
|
@@ -0,0 +1,16 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<?xml version="1.0" encoding="UTF-8"?>
|
| 2 |
+
|
| 3 |
+
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2"
|
| 4 |
+
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
|
| 5 |
+
xsi:schemaLocation="http://www.neuroml.org/schema/neuroml2 https://raw.githubusercontent.com/NeuroML/NeuroML2/master/Schemas/NeuroML2/NeuroML_v2beta3.xsd"
|
| 6 |
+
id="passiveChan">
|
| 7 |
+
|
| 8 |
+
|
| 9 |
+
<ionChannelHH id="passiveChan" conductance="10pS" type="ionChannelPassive">
|
| 10 |
+
|
| 11 |
+
<notes>Single ion channel in NeuroML2 format: passive channel providing a leak conductance </notes>
|
| 12 |
+
|
| 13 |
+
</ionChannelHH>
|
| 14 |
+
|
| 15 |
+
</neuroml>
|
| 16 |
+
|
Tutorial/Source/nmllite/report.SimHHTest.txt
ADDED
|
@@ -0,0 +1,8 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Report of running simulation with jLEMS v0.11.0
|
| 2 |
+
Simulator=jLEMS
|
| 3 |
+
SimulatorVersion=0.11.0
|
| 4 |
+
SimulationFile=/Users/padraig/neuroConstruct/osb/generic/hodgkin_huxley_tutorial/Tutorial/Source/nmllite/LEMS_SimHHTest.xml
|
| 5 |
+
StartTime=2023-11-03 11:50:42
|
| 6 |
+
SetupTime=0.241
|
| 7 |
+
RealSimulationTime=0.04
|
| 8 |
+
SimulationSaveTime=0.001
|
Tutorial/Source/passiveChan.channel.nml
ADDED
|
@@ -0,0 +1,16 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
<?xml version="1.0" encoding="UTF-8"?>
|
| 2 |
+
|
| 3 |
+
<neuroml xmlns="http://www.neuroml.org/schema/neuroml2"
|
| 4 |
+
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
|
| 5 |
+
xsi:schemaLocation="http://www.neuroml.org/schema/neuroml2 https://raw.githubusercontent.com/NeuroML/NeuroML2/master/Schemas/NeuroML2/NeuroML_v2beta3.xsd"
|
| 6 |
+
id="passiveChan">
|
| 7 |
+
|
| 8 |
+
|
| 9 |
+
<ionChannelHH id="passiveChan" conductance="10pS" type="ionChannelPassive">
|
| 10 |
+
|
| 11 |
+
<notes>Single ion channel in NeuroML2 format: passive channel providing a leak conductance </notes>
|
| 12 |
+
|
| 13 |
+
</ionChannelHH>
|
| 14 |
+
|
| 15 |
+
</neuroml>
|
| 16 |
+
|
Tutorial/Source/run.bat
ADDED
|
@@ -0,0 +1,12 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
@echo off
|
| 2 |
+
|
| 3 |
+
REM #################################################
|
| 4 |
+
REM # LEMS Hodgkin Huxley Neuron Model
|
| 5 |
+
REM #
|
| 6 |
+
REM # Command to run LEMS_HH_Simulation.xml script on Windows
|
| 7 |
+
REM #
|
| 8 |
+
REM # Usage: run.bat
|
| 9 |
+
REM #
|
| 10 |
+
REM #################################################
|
| 11 |
+
|
| 12 |
+
java -jar jNeuroML-0.7.2-jar-with-dependencies.jar LEMS_HH_Simulation.xml
|
Tutorial/Source/run.sh
ADDED
|
@@ -0,0 +1,13 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
#!/bin/bash
|
| 2 |
+
|
| 3 |
+
#################################################
|
| 4 |
+
# LEMS Hodgkin Huxley Neuron Model
|
| 5 |
+
#
|
| 6 |
+
# Command to run LEMS_HH_Simulation.xml script on Linux/Mac
|
| 7 |
+
#
|
| 8 |
+
# Usage: ./run.sh
|
| 9 |
+
#
|
| 10 |
+
#################################################
|
| 11 |
+
|
| 12 |
+
set -e
|
| 13 |
+
java -jar jNeuroML-0.7.2-jar-with-dependencies.jar LEMS_HH_Simulation.xml
|
Tutorial/Source/test/.test.jnml.omt
ADDED
|
@@ -0,0 +1,17 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Script for running automated tests on OSB using Travis-CI, see https://github.com/OpenSourceBrain/osb-model-validation
|
| 2 |
+
|
| 3 |
+
target: ../LEMS_HH_Simulation.xml
|
| 4 |
+
engine: jNeuroML
|
| 5 |
+
mep: .test.mep
|
| 6 |
+
experiments:
|
| 7 |
+
Current clamp:
|
| 8 |
+
observables:
|
| 9 |
+
spike times:
|
| 10 |
+
file:
|
| 11 |
+
path: ../hh_v.dat
|
| 12 |
+
columns: [0,1]
|
| 13 |
+
scaling: [1000, 1000]
|
| 14 |
+
spike detection:
|
| 15 |
+
method: threshold
|
| 16 |
+
threshold: 0
|
| 17 |
+
tolerance: 0.0030997162971523324
|
Tutorial/Source/test/.test.jnmlnetpyne.omt
ADDED
|
@@ -0,0 +1,17 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Script for running automated tests on OSB using Travis-CI, see https://github.com/OpenSourceBrain/osb-model-validation
|
| 2 |
+
|
| 3 |
+
target: ../LEMS_HH_Simulation.xml
|
| 4 |
+
engine: jNeuroML_NetPyNE
|
| 5 |
+
mep: .test.mep
|
| 6 |
+
experiments:
|
| 7 |
+
Current clamp:
|
| 8 |
+
observables:
|
| 9 |
+
spike times:
|
| 10 |
+
file:
|
| 11 |
+
path: ../hh_v.dat
|
| 12 |
+
columns: [0,1]
|
| 13 |
+
scaling: [1000, 1000]
|
| 14 |
+
spike detection:
|
| 15 |
+
method: threshold
|
| 16 |
+
threshold: 0
|
| 17 |
+
tolerance: 0.0
|
Tutorial/Source/test/.test.jnmlnrn.omt
ADDED
|
@@ -0,0 +1,17 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Script for running automated tests on OSB using Travis-CI, see https://github.com/OpenSourceBrain/osb-model-validation
|
| 2 |
+
|
| 3 |
+
target: ../LEMS_HH_Simulation.xml
|
| 4 |
+
engine: jNeuroML_NEURON
|
| 5 |
+
mep: .test.mep
|
| 6 |
+
experiments:
|
| 7 |
+
Current clamp:
|
| 8 |
+
observables:
|
| 9 |
+
spike times:
|
| 10 |
+
file:
|
| 11 |
+
path: ../hh_v.dat
|
| 12 |
+
columns: [0,1]
|
| 13 |
+
scaling: [1000, 1000]
|
| 14 |
+
spike detection:
|
| 15 |
+
method: threshold
|
| 16 |
+
threshold: 0
|
| 17 |
+
tolerance: 0.0
|
Tutorial/Source/test/.test.mep
ADDED
|
@@ -0,0 +1,6 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
system: Testing a single compartment cell
|
| 2 |
+
|
| 3 |
+
experiments:
|
| 4 |
+
Current clamp:
|
| 5 |
+
expected:
|
| 6 |
+
spike times: [101.94, 116.91, 131.6, 146.29, 160.97, 175.65, 190.34, 300.95, 311.36, 321.11, 330.8, 340.48, 350.15, 359.83, 369.5, 379.18, 388.86, 398.53]
|
Tutorial/Source/test/.test.python.omt
ADDED
|
@@ -0,0 +1,19 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Script for running automated tests on OSB using Travis-CI, see https://github.com/OpenSourceBrain/osb-model-validation
|
| 2 |
+
|
| 3 |
+
# Note: this is not really running with Brian2, it's just a way to get a general Python script tested on OMV...
|
| 4 |
+
|
| 5 |
+
target: ../HodgkinHuxley.py
|
| 6 |
+
engine: Brian2
|
| 7 |
+
mep: .test.mep
|
| 8 |
+
experiments:
|
| 9 |
+
Current clamp:
|
| 10 |
+
observables:
|
| 11 |
+
spike times:
|
| 12 |
+
file:
|
| 13 |
+
path: ../hh_py_v.dat
|
| 14 |
+
columns: [0,1]
|
| 15 |
+
scaling: [1, 1]
|
| 16 |
+
spike detection:
|
| 17 |
+
method: threshold
|
| 18 |
+
threshold: 0
|
| 19 |
+
tolerance: 0.001681202059472487
|
Tutorial/Source/test/.test.singleap.jnmlbrian2.omt
ADDED
|
@@ -0,0 +1,17 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Script for running automated tests on OSB using Travis-CI, see https://github.com/OpenSourceBrain/osb-model-validation
|
| 2 |
+
|
| 3 |
+
target: ../LEMS_HH_SingleAP.xml
|
| 4 |
+
engine: jNeuroML_Brian2
|
| 5 |
+
mep: .test.singleap.mep
|
| 6 |
+
experiments:
|
| 7 |
+
Current clamp:
|
| 8 |
+
observables:
|
| 9 |
+
spike times:
|
| 10 |
+
file:
|
| 11 |
+
path: ../hh_v.dat
|
| 12 |
+
columns: [0,1]
|
| 13 |
+
scaling: [1000, 1000]
|
| 14 |
+
spike detection:
|
| 15 |
+
method: threshold
|
| 16 |
+
threshold: 0
|
| 17 |
+
tolerance: 0.000375281461096
|
Tutorial/Source/test/.test.singleap.jnmleden.omt
ADDED
|
@@ -0,0 +1,17 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Script for running automated tests on OSB using Travis-CI, see https://github.com/OpenSourceBrain/osb-model-validation
|
| 2 |
+
|
| 3 |
+
target: ../LEMS_HH_SingleAP.xml
|
| 4 |
+
engine: jNeuroML_EDEN
|
| 5 |
+
mep: .test.singleap.mep
|
| 6 |
+
experiments:
|
| 7 |
+
Current clamp:
|
| 8 |
+
observables:
|
| 9 |
+
spike times:
|
| 10 |
+
file:
|
| 11 |
+
path: ../hh_v.dat
|
| 12 |
+
columns: [0,1]
|
| 13 |
+
scaling: [1000, 1000]
|
| 14 |
+
spike detection:
|
| 15 |
+
method: threshold
|
| 16 |
+
threshold: 0
|
| 17 |
+
tolerance: 1.2509382071607622e-08
|
Tutorial/Source/test/.test.singleap.jnmlmoose.omt
ADDED
|
@@ -0,0 +1,17 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Script for running automated tests on OSB using Travis-CI, see https://github.com/OpenSourceBrain/osb-model-validation
|
| 2 |
+
|
| 3 |
+
target: ../LEMS_HH_SingleAP.xml
|
| 4 |
+
engine: jNeuroML_Moose
|
| 5 |
+
mep: .test.singleap.mep
|
| 6 |
+
experiments:
|
| 7 |
+
Current clamp:
|
| 8 |
+
observables:
|
| 9 |
+
spike times:
|
| 10 |
+
file:
|
| 11 |
+
path: ../hh_v.dat
|
| 12 |
+
columns: [0,1]
|
| 13 |
+
scaling: [1000, 1000]
|
| 14 |
+
spike detection:
|
| 15 |
+
method: threshold
|
| 16 |
+
threshold: 0
|
| 17 |
+
tolerance: 0.000750562922191561
|
Tutorial/Source/test/.test.singleap.jnmlnrn.omt
ADDED
|
@@ -0,0 +1,17 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Script for running automated tests on OSB using Travis-CI, see https://github.com/OpenSourceBrain/osb-model-validation
|
| 2 |
+
|
| 3 |
+
target: ../LEMS_HH_SingleAP.xml
|
| 4 |
+
engine: jNeuroML_NEURON
|
| 5 |
+
mep: .test.singleap.mep
|
| 6 |
+
experiments:
|
| 7 |
+
Current clamp:
|
| 8 |
+
observables:
|
| 9 |
+
spike times:
|
| 10 |
+
file:
|
| 11 |
+
path: ../hh_v.dat
|
| 12 |
+
columns: [0,1]
|
| 13 |
+
scaling: [1000, 1000]
|
| 14 |
+
spike detection:
|
| 15 |
+
method: threshold
|
| 16 |
+
threshold: 0
|
| 17 |
+
tolerance: 0.0
|
Tutorial/Source/test/.test.singleap.jnmlpynnnrn.omt
ADDED
|
@@ -0,0 +1,17 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
# Script for running automated tests on OSB using Travis-CI, see https://github.com/OpenSourceBrain/osb-model-validation
|
| 2 |
+
|
| 3 |
+
target: ../LEMS_HH_SingleAP.xml
|
| 4 |
+
engine: jNeuroML_PyNN_NEURON
|
| 5 |
+
mep: .test.singleap.mep
|
| 6 |
+
experiments:
|
| 7 |
+
Current clamp:
|
| 8 |
+
observables:
|
| 9 |
+
spike times:
|
| 10 |
+
file:
|
| 11 |
+
path: ../hh_v.dat
|
| 12 |
+
columns: [0,1]
|
| 13 |
+
scaling: [1000, 1000]
|
| 14 |
+
spike detection:
|
| 15 |
+
method: threshold
|
| 16 |
+
threshold: 0
|
| 17 |
+
tolerance: 0.0
|